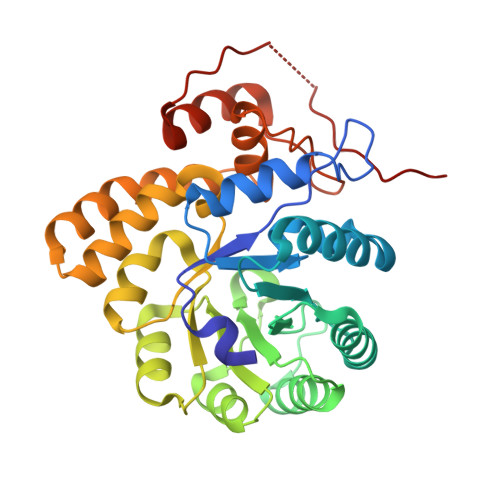
Structure, Function, Stability and Knockout Phenotype of Dihydrodipicolinate Synthase from Streptococcus pneumoniae
Dogovski, C., Gorman, M.A., Ketaren, N.E., Praszkier, J., Bryant, G., Yang, J., Griffin, M.D.W., Pearce, F.G., Bhargava, S.K., Dobson, R.C.J., Hutton, C.A., Gerrard, J.A., Robins-Browne, R.M., Jameson, G.B., Parker, M.W., Perugini, M.A.To be published.