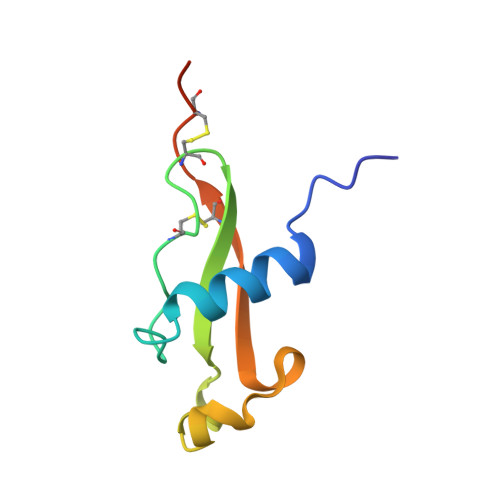
Unexpected fold in the circumsporozoite protein target of malaria vaccines.
Doud, M.B., Koksal, A.C., Mi, L.Z., Song, G., Lu, C., Springer, T.A.(2012) Proc Natl Acad Sci U S A 109: 7817-7822
- PubMed: 22547819 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1205737109
- Primary Citation Related Structures:
3VDJ, 3VDK, 3VDL - PubMed Abstract:
Circumsporozoite (CS) protein is the major surface component of Plasmodium falciparum sporozoites and is essential for host cell invasion. A vaccine containing tandem repeats, region III, and thrombospondin type-I repeat (TSR) of CS is efficacious in phase III trials but gives only a 35% reduction in severe malaria in the first year postimmunization. We solved crystal structures showing that region III and TSR fold into a single unit, an "αTSR" domain. The αTSR domain possesses a hydrophobic pocket and core, missing in TSR domains. CS binds heparin, but αTSR does not. Interestingly, polymorphic T-cell epitopes map to specialized αTSR regions. The N and C termini are unexpectedly close, providing clues for sporozoite sheath organization. Elucidation of a unique structure of a domain within CS enables rational design of next-generation subunit vaccines and functional and medicinal chemical investigation of the conserved hydrophobic pocket.
- Immune Disease Institute, Children's Hospital Boston and Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston, MA 02115, USA.
Organizational Affiliation: