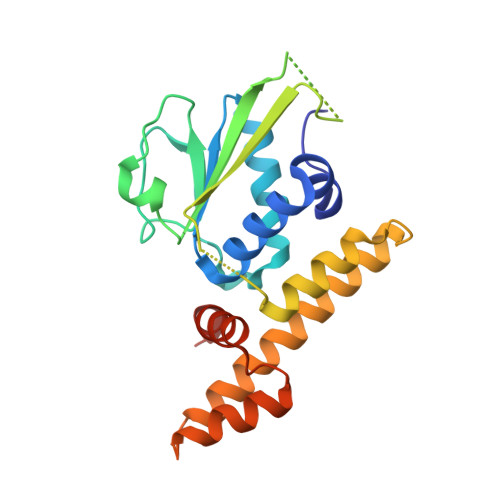
Crystal structure of folliculin reveals a hidDENN function in genetically inherited renal cancer.
Nookala, R.K., Langemeyer, L., Pacitto, A., Ochoa-Montano, B., Donaldson, J.C., Blaszczyk, B.K., Chirgadze, D.Y., Barr, F.A., Bazan, J.F., Blundell, T.L.(2012) Open Biol 2: 120071-120071
- PubMed: 22977732 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1098/rsob.120071
- Primary Citation Related Structures:
3V42 - PubMed Abstract:
Mutations in the renal tumour suppressor protein, folliculin, lead to proliferative skin lesions, lung complications and renal cell carcinoma. Folliculin has been reported to interact with AMP-activated kinase, a key component of the mammalian target of rapamycin pathway. Most cancer-causing mutations lead to a carboxy-terminal truncation of folliculin, pointing to a functional importance of this domain in tumour suppression. We present here the crystal structure of folliculin carboxy-terminal domain and demonstrate that it is distantly related to differentially expressed in normal cells and neoplasia (DENN) domain proteins, a family of Rab guanine nucleotide exchange factors (GEFs). Using biochemical analysis, we show that folliculin has GEF activity, indicating that folliculin is probably a distantly related member of this class of Rab GEFs.
- Department of Biochemistry , University of Cambridge, Sanger Building, 80 Tennis Court Road, Cambridge CB2 1GA, UK. rn229@cam.ac.uk
Organizational Affiliation: