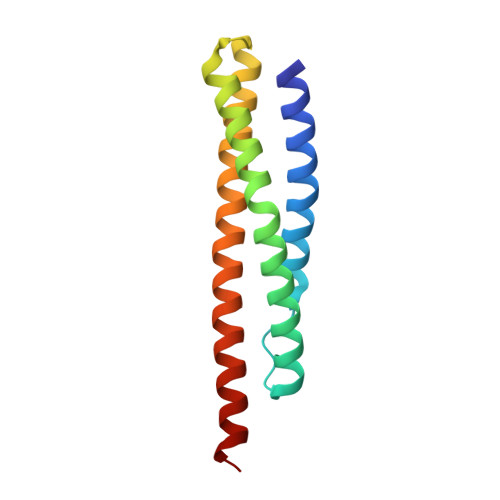
The crystal structures of dystrophin and utrophin spectrin repeats: implications for domain boundaries
Muthu, M., Richardson, K.A., Sutherland-Smith, A.J.(2012) PLoS One 7: e40066-e40066
- PubMed: 22911693 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0040066
- Primary Citation Related Structures:
3UUL, 3UUM, 3UUN - PubMed Abstract:
Dystrophin and utrophin link the F-actin cytoskeleton to the cell membrane via an associated glycoprotein complex. This functionality results from their domain organization having an N-terminal actin-binding domain followed by multiple spectrin-repeat domains and then C-terminal protein-binding motifs. Therapeutic strategies to replace defective dystrophin with utrophin in patients with Duchenne muscular dystrophy require full-characterization of both these proteins to assess their degree of structural and functional equivalence. Here the high resolution structures of the first spectrin repeats (N-terminal repeat 1) from both dystrophin and utrophin have been determined by x-ray crystallography. The repeat structures both display a three-helix bundle fold very similar to one another and to homologous domains from spectrin, α-actinin and plectin. The utrophin and dystrophin repeat structures reveal the relationship between the structural domain and the canonical spectrin repeat domain sequence motif, showing the compact structural domain of spectrin repeat one to be extended at the C-terminus relative to its previously defined sequence repeat. These structures explain previous in vitro biochemical studies in which extending dystrophin spectrin repeat domain length leads to increased protein stability. Furthermore we show that the first dystrophin and utrophin spectrin repeats have no affinity for F-actin in the absence of other domains.
- Institute of Molecular BioSciences, Massey University, Palmerston North, New Zealand.
Organizational Affiliation: