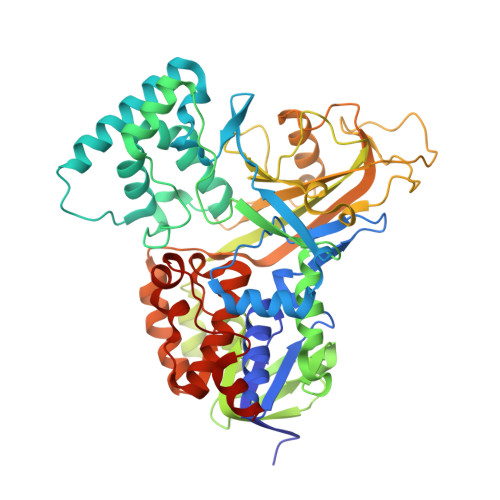
Crystal Structures and Small-angle X-ray Scattering Analysis of UDP-galactopyranose Mutase from the Pathogenic Fungus Aspergillus fumigatus.
Dhatwalia, R., Singh, H., Oppenheimer, M., Karr, D.B., Nix, J.C., Sobrado, P., Tanner, J.J.(2012) J Biological Chem 287: 9041-9051
- PubMed: 22294687 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M111.327536
- Primary Citation Related Structures:
3UTE, 3UTF, 3UTG, 3UTH - PubMed Abstract:
UDP-galactopyranose mutase (UGM) is a flavoenzyme that catalyzes the conversion of UDP-galactopyranose to UDP-galactofuranose, which is a central reaction in galactofuranose biosynthesis. Galactofuranose has never been found in humans but is an essential building block of the cell wall and extracellular matrix of many bacteria, fungi, and protozoa. The importance of UGM for the viability of many pathogens and its absence in humans make UGM a potential drug target. Here we report the first crystal structures and small-angle x-ray scattering data for UGM from the fungus Aspergillus fumigatus, the causative agent of aspergillosis. The structures reveal that Aspergillus UGM has several extra secondary and tertiary structural elements that are not found in bacterial UGMs yet are important for substrate recognition and oligomerization. Small-angle x-ray scattering data show that Aspergillus UGM forms a tetramer in solution, which is unprecedented for UGMs. The binding of UDP or the substrate induces profound conformational changes in the enzyme. Two loops on opposite sides of the active site move toward each other by over 10 Å to cover the substrate and create a closed active site. The degree of substrate-induced conformational change exceeds that of bacterial UGMs and is a direct consequence of the unique quaternary structure of Aspergillus UGM. Galactopyranose binds at the re face of the FAD isoalloxazine with the anomeric carbon atom poised for nucleophilic attack by the FAD N5 atom. The structural data provide new insight into substrate recognition and the catalytic mechanism and thus will aid inhibitor design.
- Department of Chemistry, University of Missouri, Columbia, Missouri 65211, USA.
Organizational Affiliation: