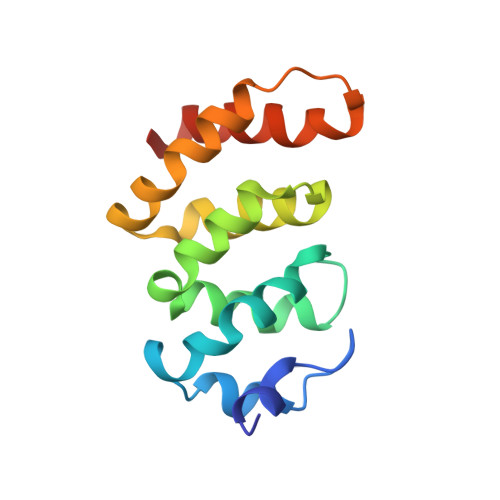
Structural view of a non pfam singleton and crystal packing analysis.
Cheng, C., Shaw, N., Zhang, X., Zhang, M., Ding, W., Wang, B.C., Liu, Z.J.(2012) PLoS One 7: e31673-e31673
- PubMed: 22363703 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0031673
- Primary Citation Related Structures:
3UT4, 3UT7, 3UT8 - PubMed Abstract:
Comparative genomic analysis has revealed that in each genome a large number of open reading frames have no homologues in other species. Such singleton genes have attracted the attention of biochemists and structural biologists as a potential untapped source of new folds. Cthe_2751 is a 15.8 kDa singleton from an anaerobic, hyperthermophile Clostridium thermocellum. To gain insights into the architecture of the protein and obtain clues about its function, we decided to solve the structure of Cthe_2751. The protein crystallized in 4 different space groups that diffracted X-rays to 2.37 Å (P3(1)21), 2.17 Å (P2(1)2(1)2(1)), 3.01 Å (P4(1)22), and 2.03 Å (C222(1)) resolution, respectively. Crystal packing analysis revealed that the 3-D packing of Cthe_2751 dimers in P4(1)22 and C222(1) is similar with only a rotational difference of 2.69° around the C axes. A new method developed to quantify the differences in packing of dimers in crystals from different space groups corroborated the findings of crystal packing analysis. Cthe_2751 is an all α-helical protein with a central hydrophobic core providing thermal stability via π:cation and π: π interactions. A ProFunc analysis retrieved a very low match with a splicing endonuclease, suggesting a role for the protein in the processing of nucleic acids. Non-Pfam singleton Cthe_2751 folds into a known all α-helical fold. The structure has increased sequence coverage of non-Pfam proteins such that more protein sequences can be amenable to modelling. Our work on crystal packing analysis provides a new method to analyze dimers of the protein crystallized in different space groups. The utility of such an analysis can be expanded to oligomeric structures of other proteins, especially receptors and signaling molecules, many of which are known to function as oligomers.
- National Laboratory of Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, Beijing, China.
Organizational Affiliation: