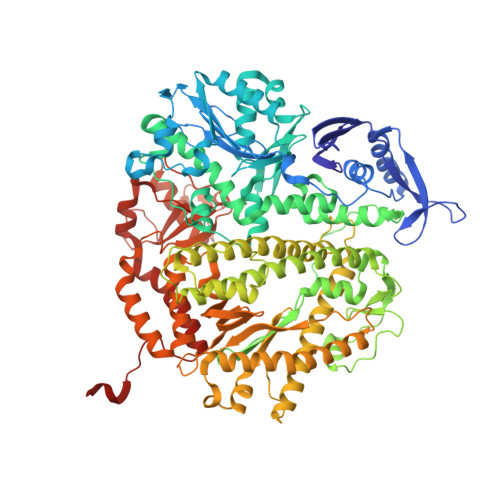
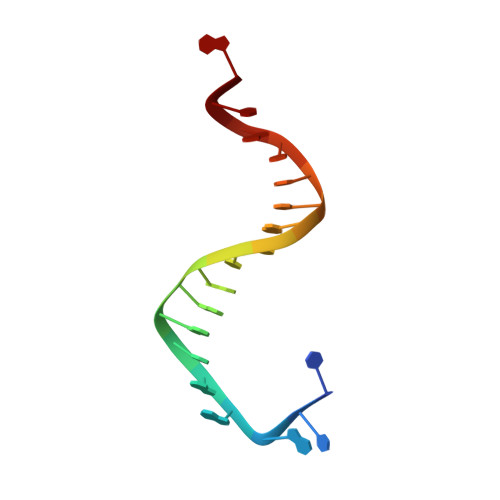
Bidentate and tridentate metal-ion coordination states within ternary complexes of RB69 DNA polymerase.
Xia, S., Eom, S.H., Konigsberg, W.H., Wang, J.(2012) Protein Sci 21: 447-451
- PubMed: 22238207 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.2026
- Primary Citation Related Structures:
3UIQ - PubMed Abstract:
Two divalent metal ions are required for primer-extension catalyzed by DNA polymerases. One metal ion brings the 3'-hydroxyl of the primer terminus and the α-phosphorus atom of incoming dNTP together for bond formation so that the catalytically relevant conformation of the triphosphate tail of the dNTP is in an α,β,γ-tridentate coordination complex with the second metal ion required for proper substrate alignment. A probable base selectivity mechanism derived from structural studies on Dpo4 suggests that the inability of mispaired dNTPs to form a substrate-aligned, tridentate coordination complex could effectively cause the mispaired dNTPs to be rejected before catalysis. Nevertheless, we found that mispaired dNTPs can actually form a properly aligned tridentate coordination complex. However, complementary dNTPs occasionally form misaligned complexes with mutant RB69 DNA polymerases (RB69pols) that are not in a tridentate coordination state. Here, we report finding a β,γ-bidentate coordination complex that contained the complementary dUpNpp opposite dA in the structure of a ternary complex formed by the wild type RB69pol at 1.88 Å resolution. Our observations suggest that several distinct metal-ion coordination states can exist at the ground state in the polymerase active site and that base selectivity is unlikely to be based on metal-ion coordination alone.
- Department of Molecular Biophysics and Biochemistry, Yale University, New Haven, Connecticut 06520-8114, USA.
Organizational Affiliation: