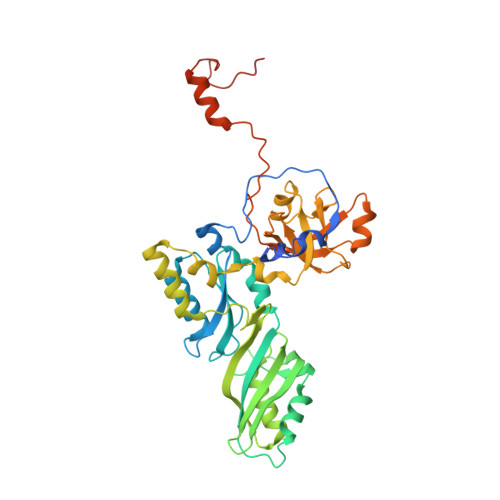
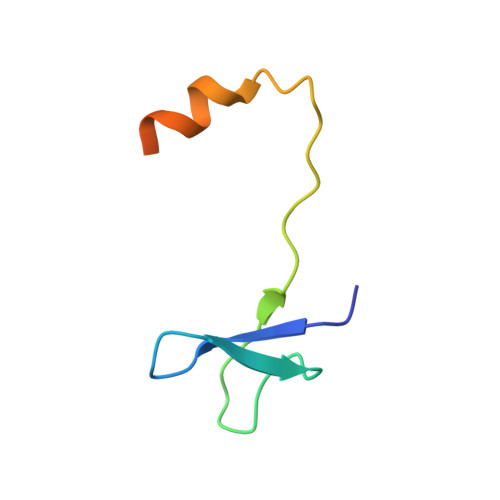
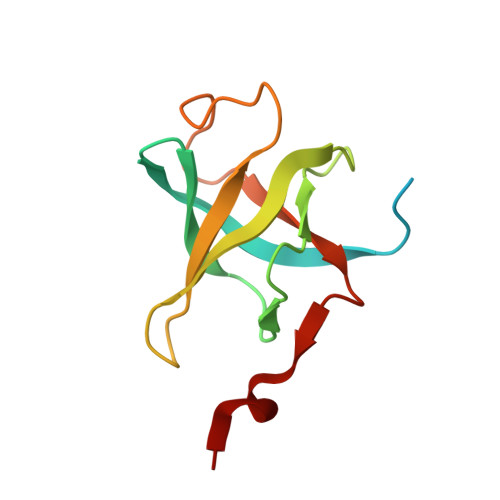
Structure of the Shq1-Cbf5-Nop10-Gar1 complex and implications for H/ACA RNP biogenesis and dyskeratosis congenita
Li, S., Duan, J., Li, D., Ma, S., Ye, K.(2011) EMBO J 30: 5010-5020
- PubMed: 22117216 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/emboj.2011.427
- Primary Citation Related Structures:
3UAH, 3UAI - PubMed Abstract:
Shq1 is a conserved protein required for the biogenesis of eukaryotic H/ACA ribonucleoproteins (RNPs), including human telomerase. We report the structure of the Shq1-specific domain alone and in complex with H/ACA RNP proteins Cbf5, Nop10 and Gar1. The Shq1-specific domain adopts a novel helical fold and primarily contacts the PUA domain and the otherwise disordered C-terminal extension (CTE) of Cbf5. The structure shows that dyskeratosis congenita mutations found in the CTE of human Cbf5 likely interfere with Shq1 binding. However, most mutations in the PUA domain are not located at the Shq1-binding surface and also have little effect on the yeast Cbf5-Shq1 interaction. Shq1 binds Cbf5 independently of the H/ACA RNP proteins Nop10, Gar1 and Nhp2 and the assembly factor Naf1, but shares an overlapping binding surface with H/ACA RNA. Shq1 point mutations that disrupt Cbf5 interaction suppress yeast growth particularly at elevated temperatures. Our results suggest that Shq1 functions as an assembly chaperone that protects the Cbf5 protein complexes from non-specific RNA binding and aggregation before assembly of H/ACA RNA.
- Department of Biochemistry and Molecular Biology, College of Life Sciences, Beijing Normal University, Beijing, China.
Organizational Affiliation: