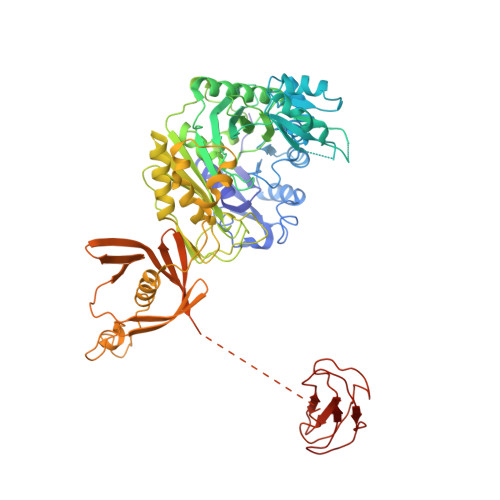
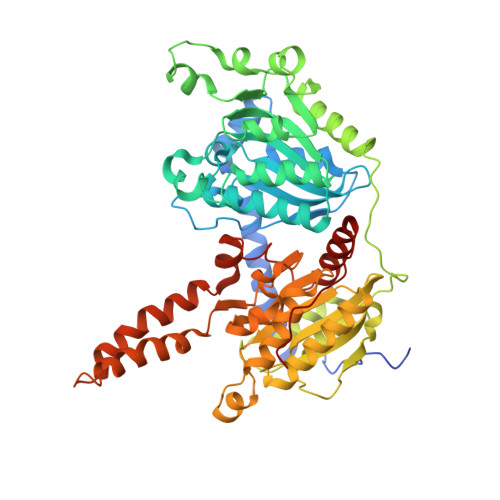
An unanticipated architecture of the 750-kDa {alpha}6{beta}6 holoenzyme of 3-methylcrotonyl-CoA carboxylase
Huang, C.S., Ge, P., Zhou, Z.H., Tong, L.(2012) Nature 481: 219-223
- PubMed: 22158123 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature10691
- Primary Citation Related Structures:
3U9R, 3U9S, 3U9T - PubMed Abstract:
3-Methylcrotonyl-CoA carboxylase (MCC), a member of the biotin-dependent carboxylase superfamily, is essential for the metabolism of leucine, and deficient mutations in this enzyme are linked to methylcrotonylglycinuria (MCG) and other serious diseases in humans. MCC has strong sequence conservation with propionyl-CoA carboxylase (PCC), and their holoenzymes are both 750-kilodalton (kDa) α(6)β(6) dodecamers. Therefore the architecture of the MCC holoenzyme is expected to be highly similar to that of PCC. Here we report the crystal structures of the Pseudomonas aeruginosa MCC (PaMCC) holoenzyme, alone and in complex with coenzyme A. Surprisingly, the structures show that the architecture and overall shape of PaMCC are markedly different when compared to PCC. The α-subunits show trimeric association in the PaMCC holoenzyme, whereas they have no contacts with each other in PCC. Moreover, the positions of the two domains in the β-subunit of PaMCC are swapped relative to those in PCC. This structural information establishes a foundation for understanding the disease-causing mutations of MCC and provides new insights into the catalytic mechanism and evolution of biotin-dependent carboxylases. The large structural differences between MCC and PCC also have general implications for the relationship between sequence conservation and structural similarity.
- Department of Biological Sciences, Columbia University, New York, New York 10027, USA.
Organizational Affiliation: