Antibacterial optimization of 4-aminothiazolyl analogues of the natural product GE2270 A: identification of the cycloalkylcarboxylic acids.
LaMarche, M.J., Leeds, J.A., Amaral, K., Brewer, J.T., Bushell, S.M., Dewhurst, J.M., Dzink-Fox, J., Gangl, E., Goldovitz, J., Jain, A., Mullin, S., Neckermann, G., Osborne, C., Palestrant, D., Patane, M.A., Rann, E.M., Sachdeva, M., Shao, J., Tiamfook, S., Whitehead, L., Yu, D.(2011) J Med Chem 54: 8099-8109
- PubMed: 21999529 Search on PubMed
- DOI: https://doi.org/10.1021/jm200938f
- Primary Citation Related Structures:
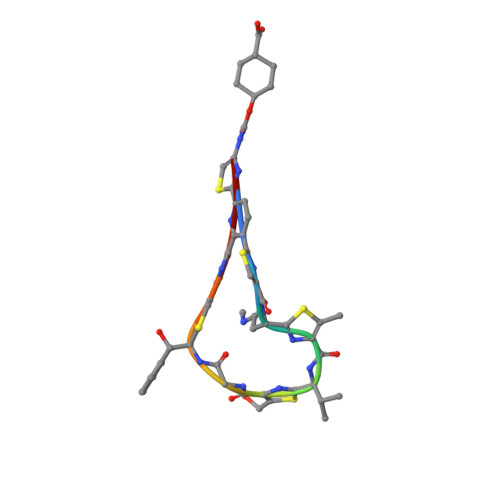
3U6B, 3U6K - PubMed Abstract:
4-Aminothiazolyl analogues of the antibiotic natural product GE2270 A (1) were designed, synthesized, and optimized for their activity against Gram positive bacterial infections. Optimization efforts focused on improving the physicochemical properties (e.g., aqueous solubility and chemical stability) of the 4-aminothiazolyl natural product template while improving the in vitro and in vivo antibacterial activity. Structure-activity relationships were defined, and the solubility and efficacy profiles were improved over those of previous analogues and 1. These studies identified novel, potent, soluble, and efficacious elongation factor-Tu inhibitors, which bear cycloalkylcarboxylic acid side chains, and culminated in the selection of development candidates amide 48 and urethane 58.
- Global Discovery Chemistry, Novartis Institutes for Biomedical Research, Cambridge, Massachusetts 02139, United States. matthew.lamarche@novartis.com
Organizational Affiliation: