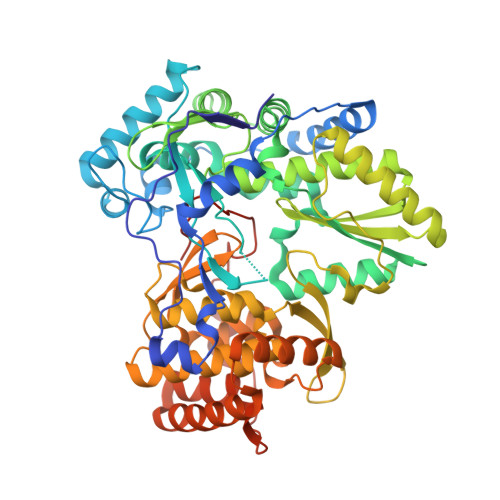
II. Novel HCV NS5B polymerase inhibitors: Discovery of indole C2 acyl sulfonamides.
Anilkumar, G.N., Selyutin, O., Rosenblum, S.B., Zeng, Q., Jiang, Y., Chan, T.Y., Pu, H., Wang, L., Bennett, F., Chen, K.X., Lesburg, C.A., Duca, J., Gavalas, S., Huang, Y., Pinto, P., Sannigrahi, M., Velazquez, F., Venkatraman, S., Vibulbhan, B., Agrawal, S., Ferrari, E., Jiang, C.K., Huang, H.C., Shih, N.Y., George Njoroge, F., Kozlowski, J.A.(2012) Bioorg Med Chem Lett 22: 713-717
- PubMed: 22104146 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2011.10.041
- Primary Citation Related Structures:
3U4O, 3U4R - PubMed Abstract:
Development of SAR at the C2 position of indole lead 1, a palm site inhibitor of HCV NS5B polymerase (NS5B IC(50)=0.053μM, replicon EC(50)=4.8μM), is described. Initial screening identified an acyl sulfonamide moiety as an isostere for the C2 carboxylic acid group. Further SAR investigation resulted in identification of acyl sufonamide analog 7q (NS5B IC(50)=0.039μM, replicon EC(50)=0.011μM) with >100-fold improved replicon activity.
- Merck Research Laboratories, 2015 Galloping Hill Road, Kenilworth, NJ 07033, USA. gopinadhan.anilkumar@Merck.com
Organizational Affiliation: