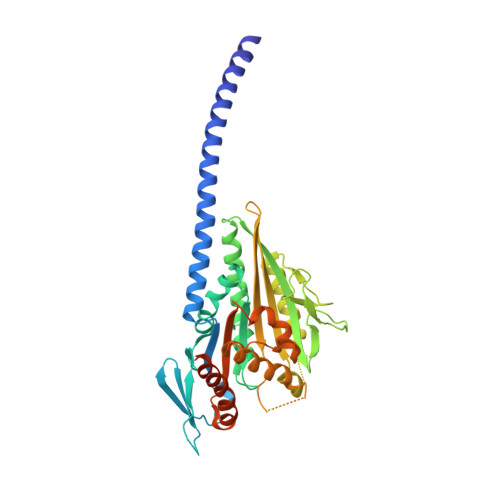
Neck-motor interactions trigger rotation of the kinesin stalk.
Liu, H.L., Pemble Iv, C.W., Endow, S.A.(2012) Sci Rep 2: 236-236
- PubMed: 22355749 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep00236
- Primary Citation Related Structures:
3U06 - PubMed Abstract:
Rotation of the coiled-coil stalk of the kinesin-14 motors is thought to drive displacements or steps by the motor along microtubules, but the structural changes that trigger stalk rotation and the nucleotide state in which it occurs are not certain. Here we report a kinesin-14 neck mutant that releases ADP more slowly than wild type and shows weaker microtubule affinity, consistent with defective stalk rotation. Unexpectedly, crystal structures show the stalk fully rotated - neck-motor interactions destabilize the stalk, causing it to rotate and ADP to be released, and alter motor affinity for microtubules. A new structural pathway accounts for the coupling of stalk rotation - the force-producing stroke - to changes in motor affinity for nucleotide and microtubules. Sequential disruption of salt bridges that stabilize the unrotated stalk could cause the stalk to initiate and complete rotation in different nucleotide states.
- Department of Cell Biology, Duke University Medical Center, Durham, NC 27710 USA.
Organizational Affiliation: