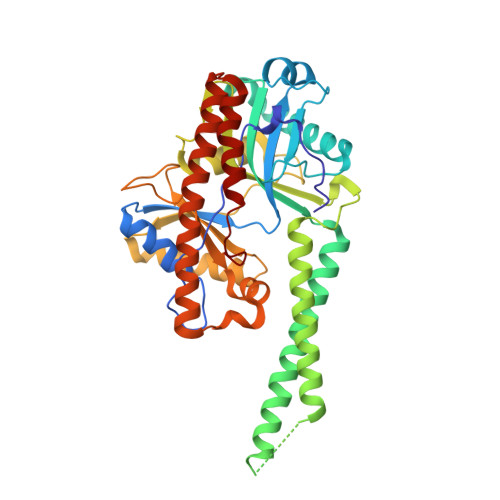
The adenylyltransferase domain of bacterial Pnkp defines a unique RNA ligase family.
Smith, P., Wang, L.K., Nair, P.A., Shuman, S.(2012) Proc Natl Acad Sci U S A 109: 2296-2301
- PubMed: 22308407 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1116827109
- Primary Citation Related Structures:
3TY5, 3TY8, 3TY9 - PubMed Abstract:
Pnkp is the end-healing and end-sealing component of an RNA repair system present in diverse bacteria from ten different phyla. To gain insight to the mechanism and evolution of this repair system, we determined the crystal structures of the ligase domain of Clostridium thermocellum Pnkp in three functional states along the reaction pathway: apoenzyme, ligase • ATP substrate complex, and covalent ligase-AMP intermediate. The tertiary structure is composed of a classical ligase nucleotidyltransferase module that is embellished by a unique α-helical insert module and a unique C-terminal α-helical module. Structure-guided mutational analysis identified active site residues essential for ligase adenylylation. Pnkp defines a new RNA ligase family with signature structural and functional properties.
- Molecular Biology Program, Sloan-Kettering Institute, 1275 York Avenue, New York, NY 10065, USA.
Organizational Affiliation: