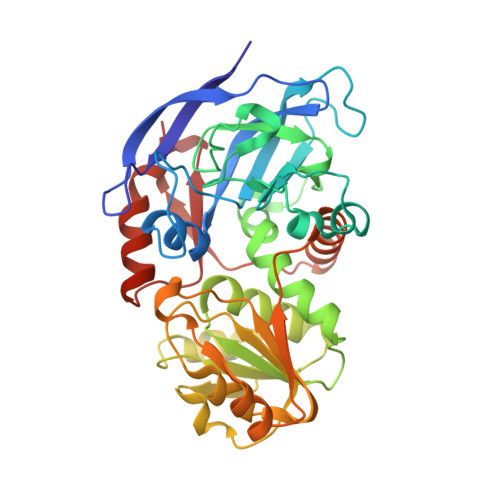
Unusual NADPH conformation in the crystal structure of a cinnamyl alcohol dehydrogenase from Helicobacter pylori in complex with NADP(H) and substrate docking analysis
Seo, K.H., Zhuang, N.N., Chen, C., Song, J.Y., Kang, H.L., Rhee, K.H., Lee, K.H.(2012) FEBS Lett 586: 337-343
- PubMed: 22269576 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2012.01.020
- Primary Citation Related Structures:
3TWO - PubMed Abstract:
Cinnamyl alcohol dehydrogenase is a zinc- and NADPH-dependent dehydrogenase catalyzing the reversible conversion of p-hydroxycinnamaldehydes to their corresponding hydroxycinnamyl alcohols. A CAD homolog from Helicobacter pylori (HpCAD) possesses broad substrate specificities like the plant CADs and additionally a dismutation activity converting benzaldehyde to benzyl alcohol and benzoic acid. We have determined the crystal structure of HpCAD complexed with NADP(H) at 2.18Å resolution to get a better understanding of this class of CAD outside of plants. The structure of HpCAD is highly homologous to the sinapyl alcohol dehydrogenase and the plant CAD with well-conserved residues involved in catalysis and zinc binding. However, the NADP(H) binding mode of the HpCAD has been found to be significantly different from those of plant CADs.
- Division of Applied Life Science (BK21 Program), Gyeongsang National University, Jinju, Republic of Korea.
Organizational Affiliation: