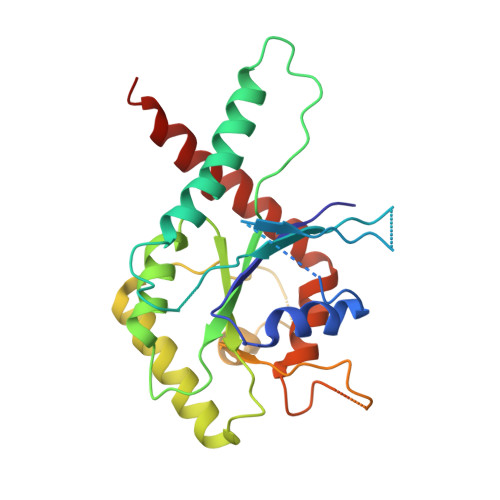
Promiscuous interactions of human septins: The GTP binding domain of SEPT7 forms filaments within the crystal.
Serrao, V.H., Alessandro, F., Caldas, V.E., Marcal, R.L., D'Muniz Pereira, H., Thiemann, O.H., Garratt, R.C.(2011) FEBS Lett 585: 3868-3873
- PubMed: 22064074 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2011.10.043
- Primary Citation Related Structures:
3TW4 - PubMed Abstract:
We describe the purification, crystallization and structure for the GTP-binding domain of human septin 7 (SEPT7G). We show that it forms filaments within the crystal lattice which employ both the G and NC interfaces, similar to those seen in the hetero-filament of SEPT2/6/7. The NC interface is considered promiscuous as it is absent from the hetero-filament. Such promiscuity could provide the potential for permuting monomers along a filament in order to generate diversity in hetero-polymers. On the other hand, our results suggest that the G and NC interfaces may be necessary but insufficient for determining correct hetero-filament assembly.
- Centro de Biotecnologia Molecular Estrutural, Instituto de Física de São Carlos, Universidade de São Paulo, São Carlos, SP, Brazil.
Organizational Affiliation: