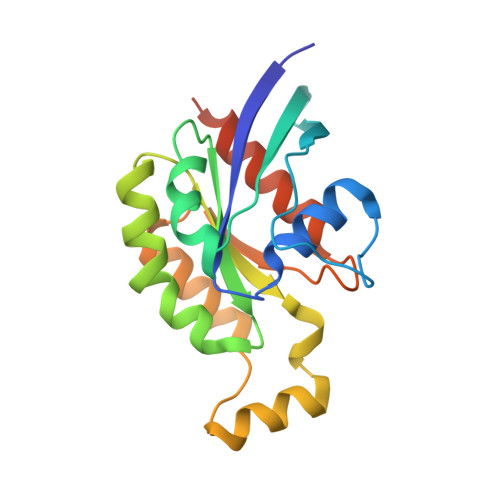
Crystal structure of mouse RhoA:GTPgammaS complex in a centered lattice.
Jobichen, C., Pal, K., Swaminathan, K.(2012) J Struct Funct Genomics 13: 241-245
- PubMed: 23001747 Search on PubMed
- DOI: https://doi.org/10.1007/s10969-012-9143-5
- Primary Citation Related Structures:
3TVD - PubMed Abstract:
RhoA, a member of the Rho sub-family of small GTPases, plays a significant signaling role in cell morphogenesis, migration, neuronal development, cell division and adhesion. So far, 4 structures of RhoA:GDP/GTP analogs and 14 structures of RhoA in complex with other proteins have been reported. All RhoA:GDP/GTP analog complexes have been crystallized in primitive lattices and RhoA is monomeric. This is the first time a RhoA:GTP analog complex has been crystallized as a dimer in a centered lattice. The present structure reveals structural differences in the switch-I (residues 28-42) and switch-II (residues 61-66) regions, which play important roles in interactions with downstream targets to transduce signals, when compared to the previously reported structures.
- Department of Biological Sciences, National University of Singapore, Singapore, Singapore.
Organizational Affiliation: