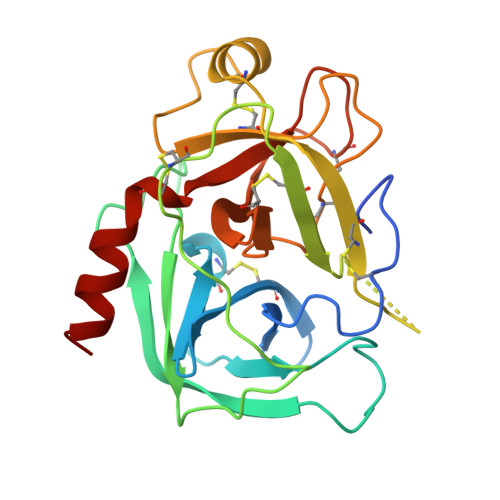
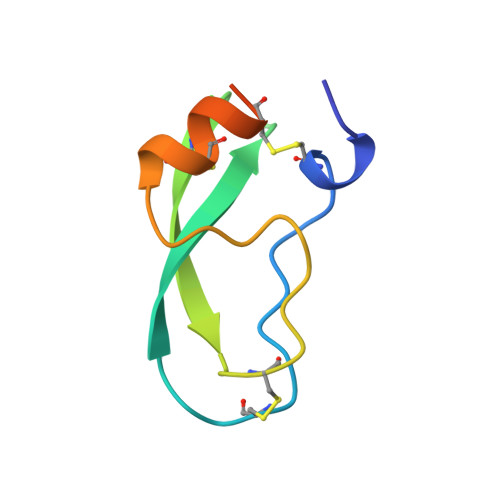
Comparison of Anionic and Cationic Trypsinogens: The Anionic Activation Domain is More Flexible in Solution and Differs in its Mode of Bpti Binding in the Crystal Structure
Pasternak, A., Ringe, D., Hedstrom, L.(1999) Protein Sci 8: 253-258
- PubMed: 10210204 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.8.1.253
- Primary Citation Related Structures:
3TGI, 3TGJ - PubMed Abstract:
Unlike bovine cationic trypsin, rat anionic trypsin retains activity at high pH. This alkaline stability has been attributed to stabilization of the salt bridge between the N-terminal Ile16 and Asp194 by the surface negative charge (Soman K, Yang A-S, Honig B, Fletterick R., 1989, Biochemistry 28:9918-9926). The formation of this salt bridge controls the conformation of the activation domain in trypsin. In this work we probe the structure of rat trypsinogen to determine the effects of the surface negative charge on the activation domain in the absence of the Ile16-Asp194 salt bridge. We determined the crystal structures of the rat trypsin-BPTI complex and the rat trypsinogen-BPTI complex at 1.8 and 2.2 A, respectively. The BPTI complex of rat trypsinogen resembles that of rat trypsin. Surprisingly, the side chain of Ile16 is found in a similar position in both the rat trypsin and trypsinogen complexes, although it is not the N-terminal residue and cannot form the salt bridge in trypsinogen. The resulting position of the activation peptide alters the conformation of the adjacent autolysis loop (residues 142-153). While bovine trypsinogen and trypsin have similar CD spectra, the CD spectrum of rat trypsinogen has only 60% of the intensity of rat trypsin. This lower intensity most likely results from increased flexibility around two conserved tryptophans, which are adjacent to the activation domain. The NMR spectrum of rat trypsinogen contains high field methyl signals as observed in bovine trypsinogen. It is concluded that the activation domain of rat trypsinogen is more flexible than that of bovine trypsinogen, but does not extend further into the protein core.
- Department of Chemistry, Brandeis University, Waltham, Massachusetts 02454, USA.
Organizational Affiliation: