Structural studies of Enterococcus faecalis methionine aminopeptidase and design of microbe specific 2,2'-bipyridine based inhibitors
Kishor, C., Gumpena, R., Reddi, R., Addlagatta, A.(2012) Medchemcomm 3: 1406-1412
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
(2012) Medchemcomm 3: 1406-1412
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Methionine aminopeptidase | 264 | Enterococcus faecalis HIP11704 | Mutation(s): 0 Gene Names: EFHG_00941, MetAP EC: 3.4.11.18 | 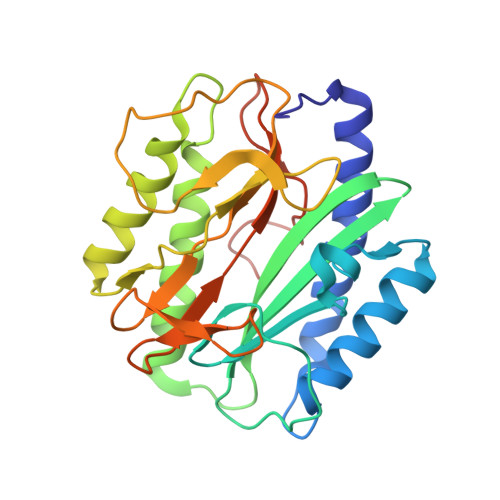 | |
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| CIT Download:Ideal Coordinates CCD File | D [auth A], E [auth B] | CITRIC ACID C6 H8 O7 KRKNYBCHXYNGOX-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 120.74 | α = 90 |
| b = 132.13 | β = 133.36 |
| c = 85.3 | γ = 90 |
| Software Name | Purpose |
|---|---|
| APEX | data collection |
| MOLREP | phasing |
| REFMAC | refinement |
| iMOSFLM | data reduction |
| SCALA | data scaling |