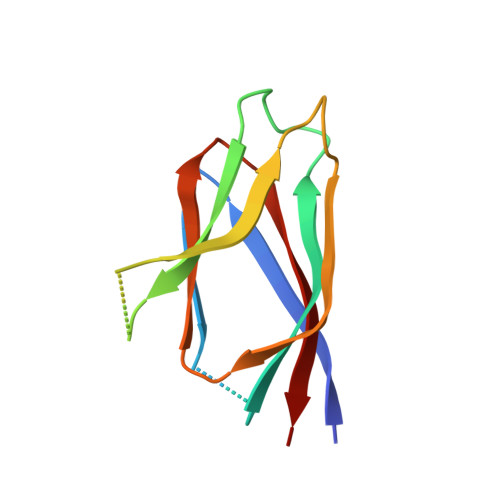
Crystal structure and function of human nucleoplasmin (npm2): a histone chaperone in oocytes and embryos.
Platonova, O., Akey, I.V., Head, J.F., Akey, C.W.(2011) Biochemistry 50: 8078-8089
- PubMed: 21863821 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi2006652
- Primary Citation Related Structures:
3T30 - PubMed Abstract:
Human Npm2 is an ortholog of Xenopus nucleoplasmin (Np), a chaperone that binds histones. We have determined the crystal structure of a truncated Npm2-core at 1.9 Å resolution and show that the N-terminal domains of Npm2 and Np form similar pentamers. This allowed us to model an Npm2 decamer which may be formed by hydrogen bonds between quasi-conserved residues in the interface between two pentamers. Interestingly, the Npm2 pentamer lacks a prototypical A1-acidic tract in each of its subunits. This feature may be responsible for the inability of Npm2-core to bind histones. However, Npm2 with a large acidic tract in its C-terminal tail (Npm2-A2) is able to bind histones and form large complexes. Fluorescence resonance energy transfer experiments and biochemical analysis of loop mutations support the premise that nucleoplasmins form decamers when they bind H2A-H2B dimers and H3-H4 tetramers simultaneously. In the absence of histone tetramers, these chaperones bind H2A-H2B dimers with a single pentamer forming the central hub. When taken together, our data provide insights into the mechanism of histone binding by nucleoplasmins.
- Department of Physiology and Biophysics, Boston University School of Medicine, 700 Albany St., Boston, Massachusetts 02118-2526, USA.
Organizational Affiliation: