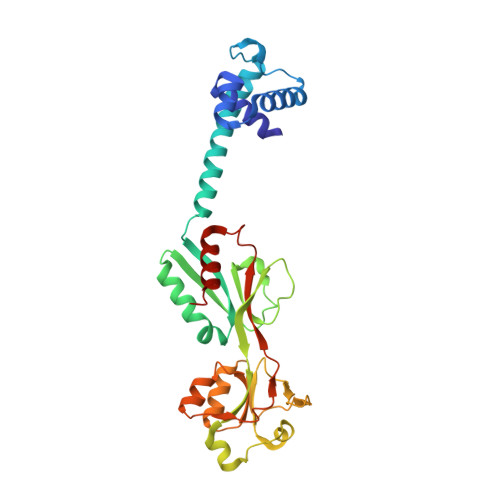
The crystal structure of AphB, a virulence gene activator from Vibrio cholerae, reveals residues that influence its response to oxygen and pH.
Taylor, J.L., De Silva, R.S., Kovacikova, G., Lin, W., Taylor, R.K., Skorupski, K., Kull, F.J.(2012) Mol Microbiol 83: 457-470
- PubMed: 22053934 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/j.1365-2958.2011.07919.x
- Primary Citation Related Structures:
3SZP, 3T1B - PubMed Abstract:
Expression of the two critical virulence factors of Vibrio cholerae, toxin-coregulated pilus and cholera toxin, is initiated at the tcpPH promoter by the regulators AphA and AphB. AphA is a winged helix DNA-binding protein that enhances the ability of AphB, a LysR-type transcriptional regulator, to activate tcpPH expression. We present here the 2.2 Å X-ray crystal structure of full-length AphB. As reported for other LysR-type proteins, AphB is a tetramer with two distinct subunit conformations. Unlike other family members, AphB must undergo a significant conformational change in order to bind to DNA. We have found five independent mutations in the putative ligand-binding pocket region that allow AphB to constitutively activate tcpPH expression at the non-permissive pH of 8.5 and in the presence of oxygen. These findings indicate that AphB is responsive to intracellular pH as well as to anaerobiosis and that residues in the ligand-binding pocket of the protein influence its ability to respond to both of these signals. We have solved the structure of one of the constitutive mutants, and observe conformational changes that would allow DNA binding. Taken together, these results describe a pathway of conformational changes allowing communication between the ligand and DNA binding regions of AphB.
- Department of Chemistry, Dartmouth College, Hanover, NH 03755, USA.
Organizational Affiliation: