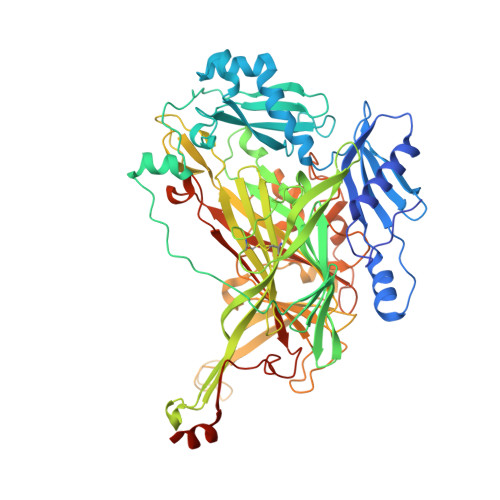
The precursor form of Hansenula polymorpha copper amine oxidase 1 in complex with CuI and CoII.
Klema, V.J., Johnson, B.J., Klinman, J.P., Wilmot, C.M.(2012) Acta Crystallogr Sect F Struct Biol Cryst Commun 68: 501-510
- PubMed: 22691777 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309112012857
- Primary Citation Related Structures:
3SX1, 3SXX, 3T0U - PubMed Abstract:
Copper amine oxidases (CAOs) catalyze the oxidative deamination of primary amines to their corresponding aldehydes, with the concomitant reduction of O(2) to H(2)O(2). Catalysis requires two cofactors: a mononuclear copper center and the cofactor 2,4,5-trihydroxyphenylalanine quinone (TPQ). TPQ is synthesized through the post-translational modification of an endogenous tyrosine residue and requires only oxygen and copper to proceed. TPQ biogenesis in CAO can be supported by alternate metals, albeit at decreased rates. A variety of factors are thought to contribute to the degree to which a metal can support TPQ biogenesis, including Lewis acidity, redox potential and electrostatic stabilization capability. The crystal structure has been solved of one of two characterized CAOs from the yeast Hansenula polymorpha (HPAO-1) in its metal-free (apo) form, which contains an unmodified precursor tyrosine residue instead of fully processed TPQ (HPAO-1 was denoted HPAO in the literature prior to 2010). Structures of apoHPAO-1 in complex with Cu(I) and Co(II) have also been solved, providing structural insight into metal binding prior to biogenesis.
- Department of Biochemistry, Molecular Biology and Biophysics, University of Minnesota, 321 Church Street SE, Minneapolis, MN 55455, USA.
Organizational Affiliation: