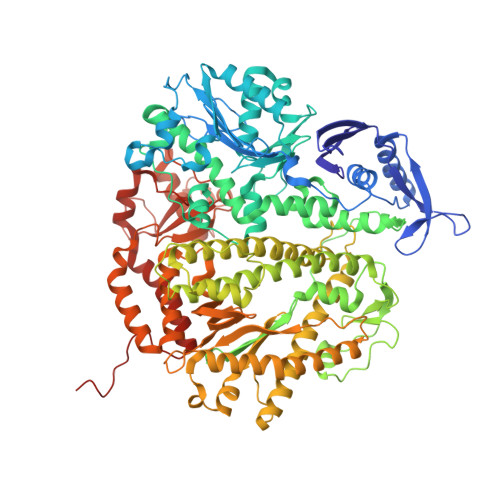
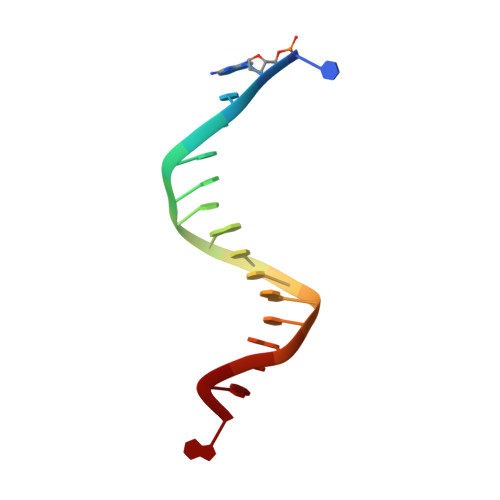
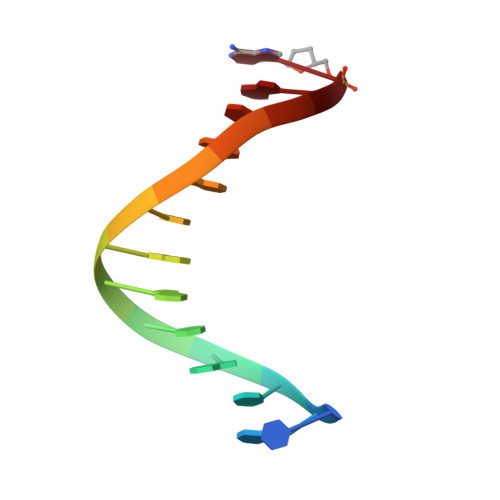
Structure of the 2-Aminopurine-Cytosine Base Pair Formed in the Polymerase Active Site of the RB69 Y567A-DNA Polymerase.
Reha-Krantz, L.J., Hariharan, C., Subuddhi, U., Xia, S., Zhao, C., Beckman, J., Christian, T., Konigsberg, W.(2011) Biochemistry 50: 10136-10149
- PubMed: 22023103 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi2014618
- Primary Citation Related Structures:
3SQ2, 3SQ4, 3SUN, 3SUO, 3SUP, 3SUQ - PubMed Abstract:
The adenine base analogue 2-aminopurine (2AP) is a potent base substitution mutagen in prokaryotes because of its enhanceed ability to form a mutagenic base pair with an incoming dCTP. Despite more than 50 years of research, the structure of the 2AP-C base pair remains unclear. We report the structure of the 2AP-dCTP base pair formed within the polymerase active site of the RB69 Y567A-DNA polymerase. A modified wobble 2AP-C base pair was detected with one H-bond between N1 of 2AP and a proton from the C4 amino group of cytosine and an apparent bifurcated H-bond between a proton on the 2-amino group of 2-aminopurine and the ring N3 and O2 atoms of cytosine. Interestingly, a primer-terminal region rich in AT base pairs, compared to GC base pairs, facilitated dCTP binding opposite template 2AP. We propose that the increased flexibility of the nucleotide binding pocket formed in the Y567A-DNA polymerase and increased "breathing" at the primer-terminal junction of A+T-rich DNA facilitate dCTP binding opposite template 2AP. Thus, interactions between DNA polymerase residues with a dynamic primer-terminal junction play a role in determining base selectivity within the polymerase active site of RB69 DNA polymerase.
- Department of Biological Sciences, University of Alberta, Edmonton, Alberta T6G 2E9, Canada. linda.reha-krantz@ualberta.ca
Organizational Affiliation: