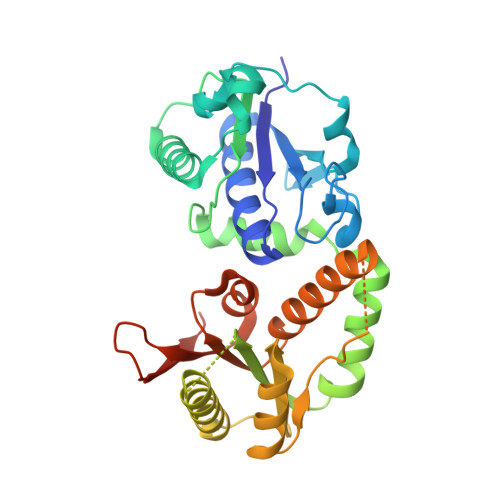
Structures of NodZ alpha-1,6-fucosyltransferase in complex with GDP and GDP-fucose
Brzezinski, K., Dauter, Z., Jaskolski, M.(2012) Acta Crystallogr D Biol Crystallogr 68: 160-168
- PubMed: 22281745 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S0907444911053157
- Primary Citation Related Structures:
3SIW, 3SIX - PubMed Abstract:
Rhizobial NodZ α1,6-fucosyltransferase (α1,6-FucT) catalyzes the transfer of the fucose (Fuc) moiety from guanosine 5'-diphosphate-β-L-fucose to the reducing end of the chitin oligosaccharide core during Nod-factor (NF) biosynthesis. NF is a key signalling molecule required for successful symbiosis with a legume host for atmospheric nitrogen fixation. To date, only two α1,6-FucT structures have been determined, both without any donor or acceptor molecule that could highlight the structural background of the catalytic mechanism. Here, the first crystal structures of α1,6-FucT in complex with its substrate GDP-Fuc and with GDP, which is a byproduct of the enzymatic reaction, are presented. The crystal of the complex with GDP-Fuc was obtained through soaking of native NodZ crystals with the ligand and its structure has been determined at 2.35 Å resolution. The fucose residue is exposed to solvent and is disordered. The enzyme-product complex crystal was obtained by cocrystallization with GDP and an acceptor molecule, penta-N-acetyl-L-glucosamine (penta-NAG). The structure has been determined at 1.98 Å resolution, showing that only the GDP molecule is present in the complex. In both structures the ligands are located in a cleft formed between the two domains of NodZ and extend towards the C-terminal domain, but their conformations differ significantly. The structures revealed that residues in three regions of the C-terminal domain, which are conserved among α1,2-, α1,6- and protein O-fucosyltransferases, are involved in interactions with the sugar-donor molecule. There is also an interaction with the side chain of Tyr45 in the N-terminal domain, which is very unusual for a GT-B-type glycosyltransferase. Only minor conformational changes of the protein backbone are observed upon ligand binding. The only exception is a movement of the loop located between strand βC2 and helix αC3. In addition, there is a shift of the αC3 helix itself upon GDP-Fuc binding.
- Synchrotron Radiation Research Section, MCL, National Cancer Institute, Argonne National Laboratory, Argonne, IL 60439, USA.
Organizational Affiliation: