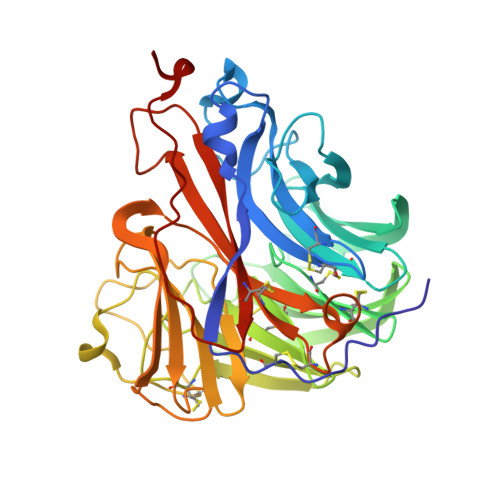
Influenza a virus n5 neuraminidase has an extended 150-cavity
Wang, M.Y., Qi, J.X., Liu, Y., Vavricka, C.J., Wu, Y., Li, Q., Gao, G.F.(2011) J Virol 85: 8431-8435
- PubMed: 21653672 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.00638-11
- Primary Citation Related Structures:
3SAL, 3SAN - PubMed Abstract:
There are 9 serotypes of neuraminidase (NA) from influenza A virus (N1 to N9), which are classified into two groups based on primary sequences (groups 1 and 2). The structural hallmark of the two groups is the presence or absence of an extra 150-cavity (formed by the 150-loop) in the active site. Thus far, structures of NAs from 6 out of the 9 serotypes have been solved. Here, we solved the N5 structure, the last unknown structure group 1 serotype with a unique Asn147 residue in its 150-loop, demonstrating that it has an extended 150-cavity that closes upon inhibitor binding.
- CAS Key Laboratory of Pathogenic Microbiology and Immunology, Institute of Microbiology, Chinese Academy of Sciences, Chaoyang District, Beijing 100101, The People's Republic of China. gaof@im.ac.cn
Organizational Affiliation: