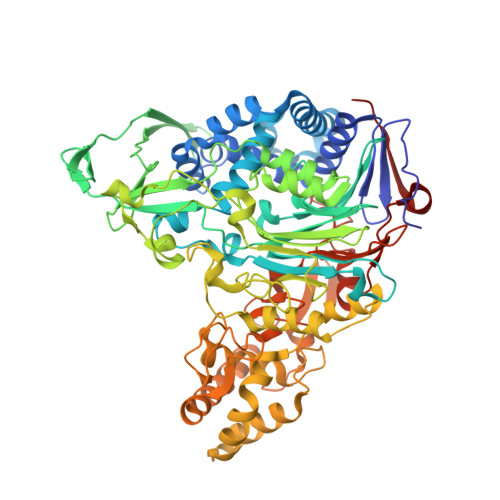
Crystal structures of glutaryl 7-aminocephalosporanic acid acylase: insight into autoproteolytic activation.
Kim, J.K., Yang, I.S., Rhee, S., Dauter, Z., Lee, Y.S., Park, S.S., Kim, K.H.(2003) Biochemistry 42: 4084-4093
- PubMed: 12680762 Search on PubMed
- DOI: https://doi.org/10.1021/bi027181x
- Primary Citation Related Structures:
1OR0, 3S8R - PubMed Abstract:
Glutaryl 7-aminocephalosporanic acid acylase (GCA, EC 3.5.1.11) is a member of N-terminal nucleophile (Ntn) hydrolases. The native enzyme is an (alpha beta)(2) heterotetramer originated from an enzymatically inactive precursor of a single polypeptide. The activation of precursor GCA consists of primary and secondary autoproteolytic cleavages, generating a terminal residue with both a nucleophile and a base and releasing a nine amino acid spacer peptide. We have determined the crystal structures of the recombinant selenomethionyl native and S170A mutant precursor from Pseudomonas sp. strain GK16. Precursor activation is likely triggered by conformational constraints within the spacer peptide, probably inducing a peptide flip. Autoproteolytic site solvent molecules, which have been trapped in a hydrophobic environment by the spacer peptide, may play a role as a general base for nucleophilic attack. The activation results in building up a catalytic triad composed of Ser170/His192/Glu624. However, the triad is not linked to the usual hydroxyl but the free alpha-amino group of the N-terminal serine residue of the native GCA. Mutagenesis and structural data support the notion that the stabilization of a transient hydroxazolidine ring during autoproteolysis would be critical during the N --> O acyl shift. The autoproteolytic activation mechanism for GCA is described.
- Graduate School of Biotechnology, Korea University, Seoul 136-701, Korea.
Organizational Affiliation: