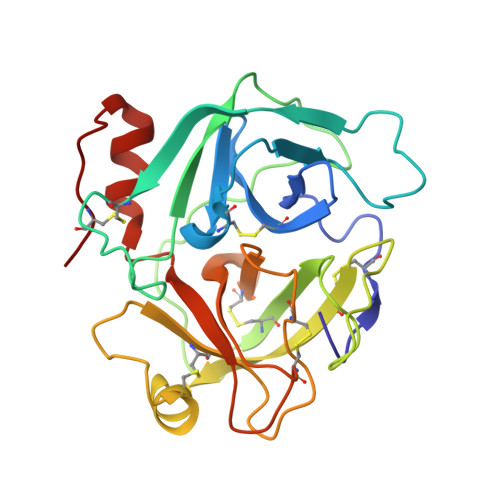
Structure of saxthrombin, a thrombin-like enzyme from Gloydius saxatilis.
Huang, K., Zhao, W., Gao, Y., Wei, W., Teng, M., Niu, L.(2011) Acta Crystallogr Sect F Struct Biol Cryst Commun 67: 862-865
- PubMed: 21821882 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309111022548
- Primary Citation Related Structures:
3S69 - PubMed Abstract:
Snake-venom thrombin-like enzymes (SVTLEs) are serine proteases that are widely distributed in snakes from the Crotalinae subfamily of the Viperidae. In contrast to other snake-venom serine proteases, they have a biochemical activity similar to that of thrombin and play an important role in the process of blood coagulation. However, SVTLEs cannot activate factor VIII, which is essential in blood-clot stabilization. Consequently, blood clots produced by SVTLEs are not stable and are cleared rapidly. This characteristic makes SVTLEs attractive as potential candidates for antithrombotic therapy. Saxthrombin, an SVTLE from Gloydius saxatilis, was purified and crystallized to obtain a high-quality crystal, from which data were acquired to 1.43 Å resolution. Preliminary X-ray diffraction analysis showed that the crystal belonged to space group C2, with unit-cell parameters a = 94.2, b = 52.2, c = 50.1 Å, β = 96.7°. The crystal structure was determined by molecular replacement and the final R factor was 18.69%; the R(free) was 20.01%. This is the first report of a crystal structure of an SVTLE. Saxthrombin belongs to the typical α/β-hydrolase fold of serine proteases. Its structure was compared with those of thrombin and other snake-venom serine proteases. The observed differences in the amino-acid composition of the loops surrounding the active site appear to contribute to different surface-charge distributions and thus alter the shape of the active-site cleft, which may explain the differences in substrate affinity.
- Hefei National Laboratory for Physical Sciences at Microscale and School of Life Sciences, University of Science and Technology of China, Hefei, Anhui, People's Republic of China.
Organizational Affiliation: