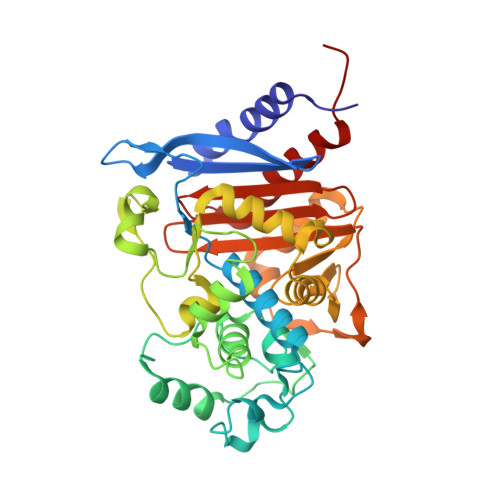
Structural analysis of the Asn152Gly mutant of P99 cephalosporinase.
Ruble, J.F., Lefurgy, S.T., Cornish, V.W., Powers, R.A.(2012) Acta Crystallogr D Biol Crystallogr 68: 1189-1193
- PubMed: 22948919 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444912024080
- Primary Citation Related Structures:
3S4X - PubMed Abstract:
P99 cephalosporinase is a class C β-lactamase that is responsible in part for the widespread bacterial resistance to β-lactam antibiotics. Mutations of the conserved active-site residue Asn152 of the enzyme have been shown to alter β-lactam substrate specificity in vivo. Mutation of Asn152 to a glycine is notable in that it exhibits in vivo substrate-selectivity switching. In order to better understand the structural basis for this observed switch, the X-ray crystal structure of the apo Asn152Gly mutant of P99 was determined to 1.95 Å resolution. Unexpectedly, the artificial C-terminal His(6) tag of a symmetrically-related molecule was observed bound in the active site. The His(6) tag makes several interactions with key active-site residues, as well as with several sulfate ions. Additionally, the overall C-terminus occupies the space left vacant upon the mutation of Asn152 to glycine.
- Department of Chemistry, Grand Valley State University, Allendale, MI 49401, USA.
Organizational Affiliation: