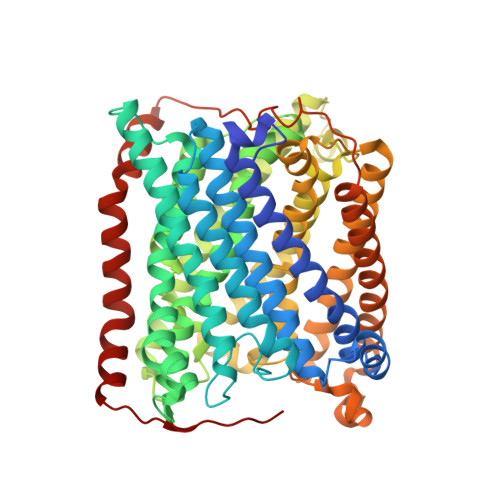
Mobility of Xe atoms within the oxygen diffusion channel of cytochrome ba(3) oxidase.
Luna, V.M., Fee, J.A., Deniz, A.A., Stout, C.D.(2012) Biochemistry 51: 4669-4676
- PubMed: 22607023 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi3003988
- Primary Citation Related Structures:
3S33, 3S38, 3S39, 3S3A, 3S3B, 3S3C, 3S3D - PubMed Abstract:
We use a form of "freeze-trap, kinetic crystallography" to explore the migration of Xe atoms away from the dinuclear heme a(3)/Cu(B) center in Thermus thermophilus cytochrome ba(3) oxidase. This enzyme is a member of the heme-copper oxidase superfamily and is thus crucial for dioxygen-dependent life. The mechanisms involved in the migration of oxygen, water, electrons, and protons into and/or out of the specialized channels of the heme-copper oxidases are generally not well understood. Pressurization of crystals with Xe gas previously revealed a O(2) diffusion channel in cytochrome ba(3) oxidase that is continuous, Y-shaped, 18-20 Å in length and comprised of hydrophobic residues, connecting the protein surface within the bilayer to the a(3)-Cu(B) center in the active site. To understand movement of gas molecules within the O(2) channel, we performed crystallographic analysis of 19 Xe laden crystals freeze-trapped in liquid nitrogen at selected times between 0 and 480 s while undergoing outgassing at room temperature. Variation in Xe crystallographic occupancy at five discrete sites as a function of time leads to a kinetic model revealing relative degrees of mobility of Xe atoms within the channel. Xe egress occurs primarily through the channel formed by the Xe1 → Xe5 → Xe3 → Xe4 sites, suggesting that ingress of O(2) is likely to occur by the reverse of this process. The channel itself appears not to undergo significant structural changes during Xe migration, thereby indicating a passive role in this important physiological function.
- Department of Molecular Biology, The Scripps Research Institute, La Jolla, CA 92037, USA. vmitchluna@gmail.com
Organizational Affiliation: