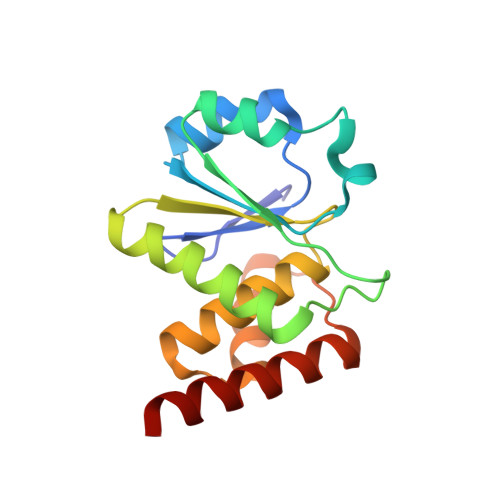
Structural and functional analysis of PTPMT1, a phosphatase required for cardiolipin synthesis.
Xiao, J., Engel, J.L., Zhang, J., Chen, M.J., Manning, G., Dixon, J.E.(2011) Proc Natl Acad Sci U S A 108: 11860-11865
- PubMed: 21730175 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1109290108
- Primary Citation Related Structures:
3RGO, 3RGQ - PubMed Abstract:
PTPMT1 (PTP localized to the Mitochondrion 1) is a member of the protein tyrosine phosphatase superfamily that is localized exclusively to the mitochondrion. We recently reported that PTPMT1 dephosphorylates phosphatidylglycerol phosphate, an essential intermediate of cardiolipin biosynthesis. To gain further insights into the molecular basis of PTPMT1 function, we determined the crystal structures of the phosphatase domain of PTPMT1. PTPMT1 exhibits a canonical protein tyrosine phosphatase domain fold, resembling many dual-specificity phosphatases such as phosphatase and tensin homolog and vaccinia H1-related phosphatase. We also determined the structure of the catalytically inactive phosphatase in complex with a surrogate substrate, phosphatidylinositol 5-phosphate, which sheds light on the substrate recognition and specificity of PTPMT1. Comparison of the apo and substrate-bound structures of PTPMT1 suggests that it undergoes significant conformational change during catalysis, and we further demonstrated that an evolutionarily conserved EEYE loop is important for its activity.
- Department of Pharmacology, University of California, La Jolla, CA 92093, USA.
Organizational Affiliation: