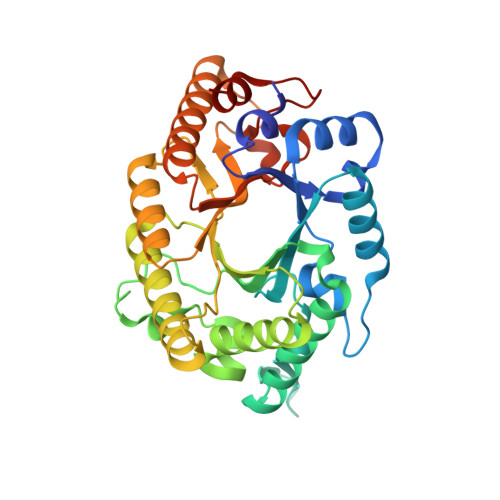
Novel structural features of xylanase A1 from Paenibacillus sp. JDR-2.
St John, F.J., Preston, J.F., Pozharski, E.(2012) J Struct Biol 180: 303-311
- PubMed: 23000703 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2012.09.007
- Primary Citation Related Structures:
3RDK, 3RO8, 4E4P - PubMed Abstract:
The Gram-positive bacterium Paenibacillus sp. JDR-2 (PbJDR2) has been shown to have novel properties in the utilization of the abundant but chemically complex hemicellulosic sugar glucuronoxylan. Xylanase A1 of PbJDR2 (PbXynA1) has been implicated in an efficient process in which extracellular depolymerization of this polysaccharide is coupled to assimilation and intracellular metabolism. PbXynA1is a 154kDa cell wall anchored multimodular glycosyl hydrolase family 10 (GH10) xylanase. In this work, the 38kDa catalytic module of PbXynA1 has been structurally characterized revealing several new features not previously observed in structures of GH10 xylanases. These features are thought to facilitate hydrolysis of highly substituted, chemically complex xylans that may be the form found in close proximity to the cell wall of PbJDR2, an organism shown to have a preference for growth on polymeric glucuronoxylan.
- Department of Pharmaceutical Sciences, University of Maryland School of Pharmacy, 20 Penn Street, Baltimore, MD 21201, USA. fjstjohn@gmail.com
Organizational Affiliation: