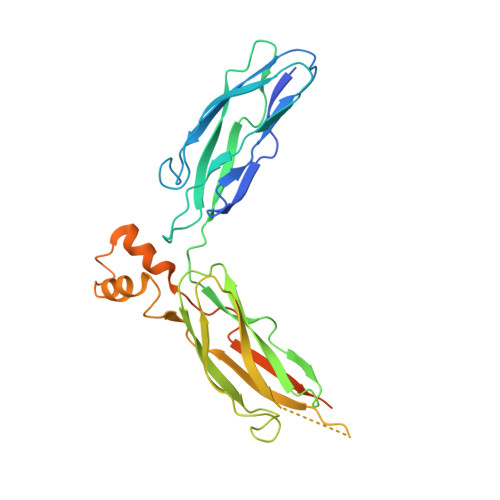
Structural Basis of the Ca(2+) Inhibitory Mechanism of Drosophila Na(+)/Ca(2+) Exchanger CALX and Its Modification by Alternative Splicing.
Wu, M., Tong, S., Gonzalez, J., Jayaraman, V., Spudich, J.L., Zheng, L.(2011) Structure 19: 1509-1517
- PubMed: 22000518 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2011.07.008
- Primary Citation Related Structures:
3RB5, 3RB7 - PubMed Abstract:
The Na(+)/Ca(2+) exchanger CALX promotes Ca(2+) efflux in Drosophila sensory neuronal cells to facilitate light-mediated Ca(2+) homeostasis. CALX activity is negatively regulated by specific Ca(2+) interaction within its two intracellular Ca(2+) regulatory domains CBD1 and CBD2, yet how the Ca(2+) binding is converted to molecular motion to operate the exchanger is unknown. Here, we report crystal structures of the entire Ca(2+) regulatory domain CBD12 from two alternative splicing isoforms, CALX 1.1 and 1.2, exhibiting distinct regulatory Ca(2+) dependency. The structures show an open V-shaped conformation with four Ca(2+) ions bound on the CBD domain interface, confirmed by LRET analysis. The structures together with Ca(2+)-binding analysis support that the Ca(2+) inhibition of CALX is achieved by interdomain conformational changes induced by Ca(2+) binding at CBD1. The conformational difference between the two isoforms also indicates that alternative splicing adjusts the interdomain orientation angle to modify the Ca(2+) regulatory property of the exchangers.
- Center for Membrane Biology, Department of Biochemistry and Molecular Biology, The University of Texas Houston Medical School, Houston, TX 77030, USA.
Organizational Affiliation: