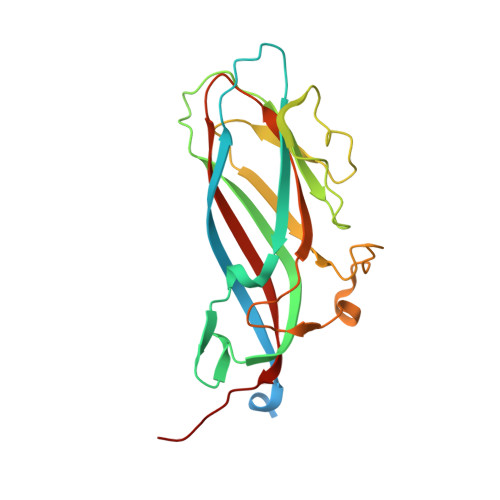
The 2.3-angstrom structure of porcine circovirus 2.
Khayat, R., Brunn, N., Speir, J.A., Hardham, J.M., Ankenbauer, R.G., Schneemann, A., Johnson, J.E.(2011) J Virol 85: 7856-7862
- PubMed: 21632760 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.00737-11
- Primary Citation Related Structures:
3R0R - PubMed Abstract:
Porcine circovirus 2 (PCV2) is a T=1 nonenveloped icosahedral virus that has had severe impact on the swine industry. Here we report the crystal structure of an N-terminally truncated PCV2 virus-like particle at 2.3-Å resolution, and the cryo-electron microscopy (cryo-EM) image reconstruction of a full-length PCV2 virus-like particle at 9.6-Å resolution. This is the first atomic structure of a circovirus. The crystal structure revealed that the capsid protein fold is a canonical viral jelly roll. The loops connecting the strands of the jelly roll define the limited features of the surface. Sulfate ions interacting with the surface and electrostatic potential calculations strongly suggest a heparan sulfate binding site that allows PCV2 to gain entry into the cell. The crystal structure also allowed previously determined epitopes of the capsid to be visualized. The cryo-EM image reconstruction showed that the location of the N terminus, absent in the crystal structure, is inside the capsid. As the N terminus was previously shown to be antigenic, it may externalize through viral "breathing."
- Department of Molecular Biology, The Scripps Research Institute, La Jolla, CA 92037, USA. rkhayat@scripps.edu
Organizational Affiliation: