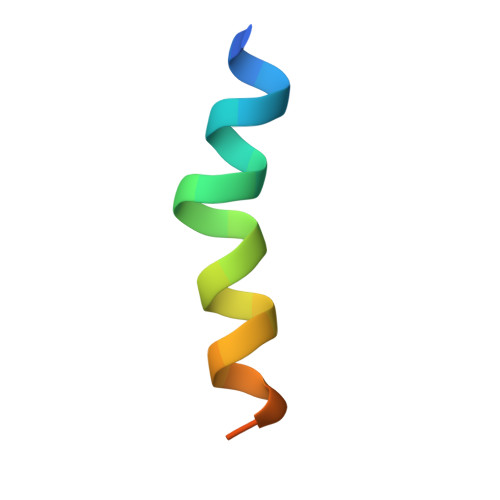
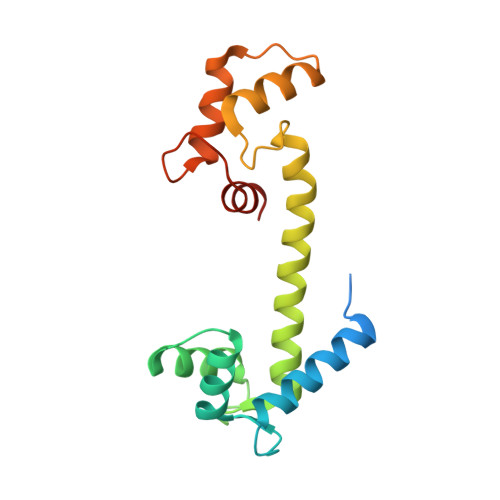
The structure, molecular dynamics, and energetics of centrin-melittin complex.
Sosa Ldel, V., Alfaro, E., Santiago, J., Narvaez, D., Rosado, M.C., Rodriguez, A., Gomez, A.M., Schreiter, E.R., Pastrana-Rios, B.(2011) Proteins 79: 3132-3143
- PubMed: 21989934 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/prot.23142
- Primary Citation Related Structures:
3QRX - PubMed Abstract:
Centrin is a calcium binding protein (CaBP) belonging to the EF-hand superfamily. As with other proteins within this family, centrin is a calcium sensor with multiple biological target proteins. We chose to study Chlamydomonas reinhardtii centrin (Crcen) and its interaction with melittin (MLT) as a model for CaBP complexes due to its amphipathic properties. Our goal was to determine the molecular interactions that lead to centrin-MLT complex formation, their relative stability, and the conformational changes associated with the interaction, when compared to the single components. For this, we determined the thermodynamic parameters that define Crcen-MLT complex formation. Two-dimensional infrared (2D IR) correlation spectroscopy were used to study the amide I', I'*, and side chain bands for (13)C-Crcen, MLT, and the (13)C-Crcen-MLT complex. This approach resulted in the determination of MLT's increased helicity, while centrin was stabilized within the complex. Herein we provide the first complete molecular description of centrin-MLT complex formation and the dissociation process. Also, discussed is the first structure of a CaBP-MLT complex by X-ray crystallography, which shows that MLT has a different binding orientation than previously characterized centrin-bound peptides. Finally, all of the experimental results presented herein are consistent with centrin maintaining an extended conformation while interacting with MLT. The molecular implications of these results are: (1) the recognition of hydrophobic contacts as requirements for initial binding, (2) minimum electrostatic interactions within the C-terminal end of the peptide, and (3) van der Waals interactions within MLTs N-terminal end are required for complex formation.
- Department of Chemistry, University of Puerto Rico, Mayagüez, Puerto Rico 00681-9019.
Organizational Affiliation: