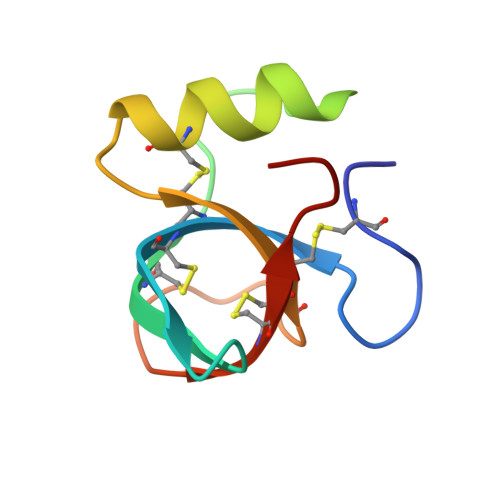
Amphiphilic nanotubes in the crystal structure of a biosurfactant protein hydrophobin HFBII.
Kallio, J.M., Rouvinen, J.(2011) Chem Commun (Camb) 47: 9843-9845
- PubMed: 21808803 Search on PubMed
- DOI: https://doi.org/10.1039/c1cc13139g
- Primary Citation Related Structures:
3QQT - PubMed Abstract:
Atomic scale experimental data by X-ray crystallography have been collected on an amphiphilic protein nanotube, consisting of a biosurfactant protein Trichoderma reesei hydrophobin HFBII.
- Department of Chemistry, University of Eastern Finland, Joensuu, Finland. johanna.kallio@embl-hamburg.de
Organizational Affiliation: