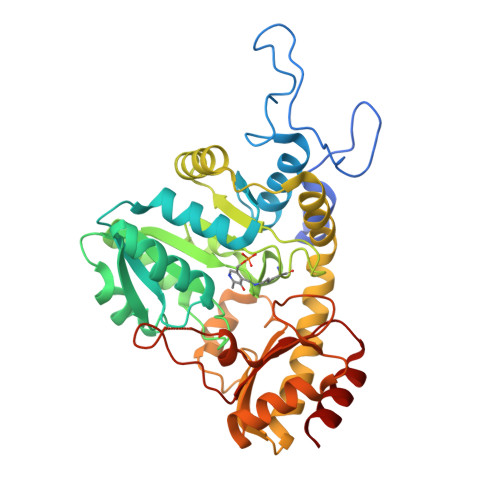
Structure of the cystathionine [gamma]-synthase MetB from Mycobacterium ulcerans
Clifton, M.C., Abendroth, J., Edwards, T.E., Leibly, D.J., Gillespie, A.K., Ferrell, M., Dieterich, S.H., Exley, I., Staker, B.L., Myler, P.J., Van Voorhis, W.C., Stewart, L.J.(2011) Acta Crystallogr Sect F Struct Biol Cryst Commun 67: 1154-1158
- PubMed: 21904066 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309111029575
- Primary Citation Related Structures:
3QHX, 3QI6 - PubMed Abstract:
Cystathionine γ-synthase (CGS) is a transulfurication enzyme that catalyzes the first specific step in L-methionine biosynthesis by the reaction of O(4)-succinyl-L-homoserine and L-cysteine to produce L-cystathionine and succinate. Controlling the first step in L-methionine biosythesis, CGS is an excellent potential drug target. Mycobacterium ulcerans is a slow-growing mycobacterium that is the third most common form of mycobacterial infection, mainly infecting people in Africa, Australia and Southeast Asia. Infected patients display a variety of skin ailments ranging from indolent non-ulcerated lesions as well as ulcerated lesions. Here, the crystal structure of CGS from M. ulcerans covalently linked to the cofactor pyridoxal phosphate (PLP) is reported at 1.9 Å resolution. A second structure contains PLP as well as a highly ordered HEPES molecule in the active site acting as a pseudo-ligand. These results present the first structure of a CGS from a mycobacterium and allow comparison with other CGS enzymes. This is also the first structure reported from the pathogen M. ulcerans.
- Seattle Structural Genomics Center for Infectious Disease (SSGCID), USA. mclifton@embios.com
Organizational Affiliation: