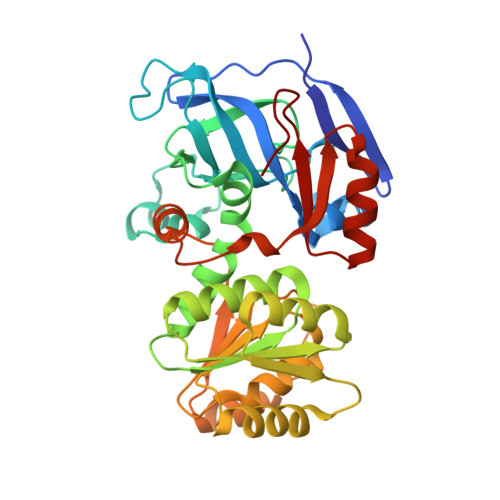
X-ray crystal structure and small-angle X-ray scattering of sheep liver sorbitol dehydrogenase.
Yennawar, H., Moller, M., Gillilan, R., Yennawar, N.(2011) Acta Crystallogr D Biol Crystallogr 67: 440-446
- PubMed: 21543846 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S0907444911007815
- Primary Citation Related Structures:
3QE3 - PubMed Abstract:
The X-ray crystal structure of sheep liver sorbitol dehydrogenase (slSDH) has been determined using the crystal structure of human sorbitol dehydrogenase (hSDH) as a molecular-replacement model. slSDH crystallized in space group I222 with one monomer in the asymmetric unit. A conserved tetramer that superposes well with that seen in hSDH (despite belonging to a different space group) and obeying the 222 crystal symmetry is seen in slSDH. An acetate molecule is bound in the active site, coordinating to the active-site zinc through a water molecule. Glycerol, a substrate of slSDH, also occupies the substrate-binding pocket together with the acetate designed by nature to fit large polyol substrates. The substrate-binding pocket is seen to be in close proximity to the tetramer interface, which explains the need for the structural integrity of the tetramer for enzyme activity. Small-angle X-ray scattering was also used to identify the quaternary structure of the tetramer of slSDH in solution.
- Department of Biochemistry and Molecular Biology and Huck Institutes of Life Sciences, Pennsylvania State University, 8 Althouse Laboratory, University Park, PA 16802, USA.
Organizational Affiliation: