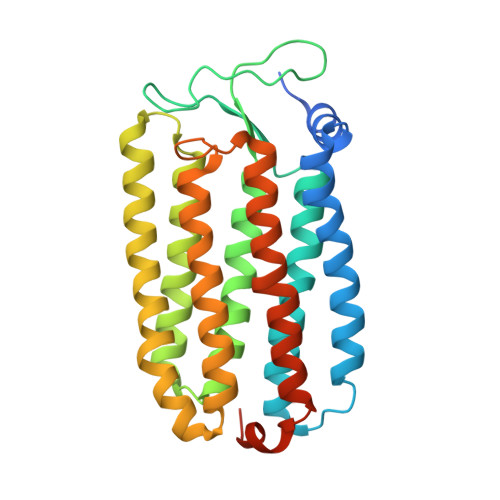
Crystal structures of an O-like blue form and an anion-free yellow form of pharaonis halorhodopsin
Kanada, S., Takeguchi, Y., Murakami, M., Ihara, K., Kouyama, T.(2011) J Mol Biology 413: 162-176
- PubMed: 21871461 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2011.08.021
- Primary Citation Related Structures:
3QBG, 3QBI, 3QBK - PubMed Abstract:
Halorhodopsin from Natronomonas pharaonis (pHR) was previously crystallized into a monoclinic space group C2, and the structure of the chloride-bound purple form was determined. Here, we report the crystal structures of two chloride-free forms of pHR, that is, an O-like blue form and an M-like yellow form. When the C2 crystal was soaked in a chloride-free alkaline solution, the protein packing was largely altered and the yellow form containing all-trans retinal was generated. Upon neutralization, this yellow form was converted into the blue form. From structural comparison of the different forms of pHR, it was shown that the removal of a chloride ion from the primary binding site (site I), which is located between the retinal Schiff base and Thr126, is accompanied by such a deformation of helix C that the side chain of Thr126 moves toward helix G, leading to a significant shrinkage of site I. A large structural change is also induced in the chloride uptake pathway, where a flip motion of the side chain of Glu234 is accompanied by large movements of the surrounding aromatic residues. Irrespective of different charge distributions at the active site, there was no large difference in the structures of the yellow form and the blue form. It is shown that the yellow-to-purple transition is initiated by the entrance of one water and one HCl to the active site, where the proton and the chloride ion in HCl are transferred to the Schiff base and site I, respectively.
- Department of Physics, Graduate School of Science, Nagoya University, Nagoya 464-8602, Japan.
Organizational Affiliation: