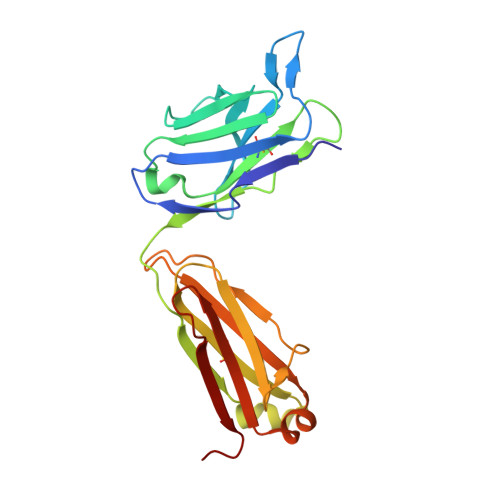
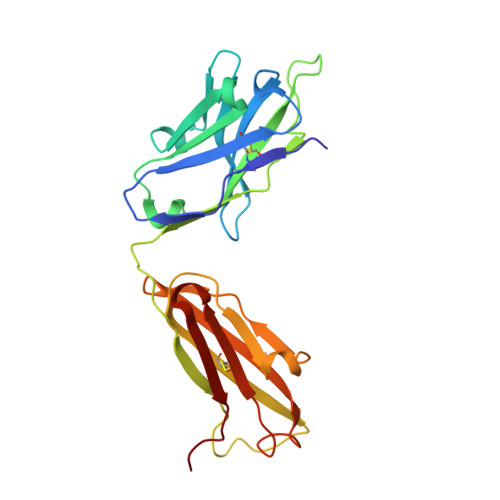
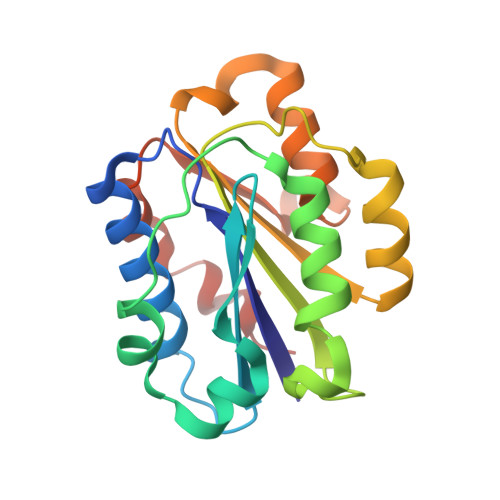
Stable Coordination of the Inhibitory Ca2+ Ion at the Metal Ion-Dependent Adhesion Site in Integrin CD11b/CD18 by an Antibody-Derived Ligand Aspartate: Implications for Integrin Regulation and Structure-Based Drug Design.
Mahalingam, B., Ajroud, K., Alonso, J.L., Anand, S., Adair, B.D., Horenstein, A.L., Malavasi, F., Xiong, J.P., Arnaout, M.A.(2011) J Immunol 187: 6393-6401
- PubMed: 22095715 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.4049/jimmunol.1102394
- Primary Citation Related Structures:
3Q3G, 3QA3 - PubMed Abstract:
A central feature of integrin interaction with physiologic ligands is the monodentate binding of a ligand carboxylate to a Mg(2+) ion hexacoordinated at the metal ion-dependent adhesion site (MIDAS) in the integrin A domain. This interaction stabilizes the A domain in the high-affinity state, which is distinguished from the default low-affinity state by tertiary changes in the domain that culminate in cell adhesion. Small molecule ligand-mimetic integrin antagonists act as partial agonists, eliciting similar activating conformational changes in the A domain, which has contributed to paradoxical adhesion and increased patient mortality in large clinical trials. As with other ligand-mimetic integrin antagonists, the function-blocking mAb 107 binds MIDAS of integrin CD11b/CD18 A domain (CD11bA), but in contrast, it favors the inhibitory Ca(2+) ion over the Mg(2+) ion at MIDAS. We determined the crystal structures of the Fab fragment of mAb 107 complexed to the low- and high-affinity states of CD11bA. Favored binding of the Ca(2+) ion at MIDAS is caused by the unusual symmetric bidentate ligation of a Fab-derived ligand Asp to a heptacoordinated MIDAS Ca(2+) ion. Binding of the Fab fragment of mAb 107 to CD11bA did not trigger the activating tertiary changes in the domain or in the full-length integrin. These data show that the denticity of the ligand Asp/Glu can modify the divalent cation selectivity at MIDAS and hence integrin function. Stabilizing the Ca(2+) ion at MIDAS by bidentate ligation to a ligand Asp/Glu may provide one approach for designing pure integrin antagonists.
- Structural Biology Program, Division of Nephrology, Massachusetts General Hospital, Harvard Medical School, Charlestown, MA 02129, USA.
Organizational Affiliation: