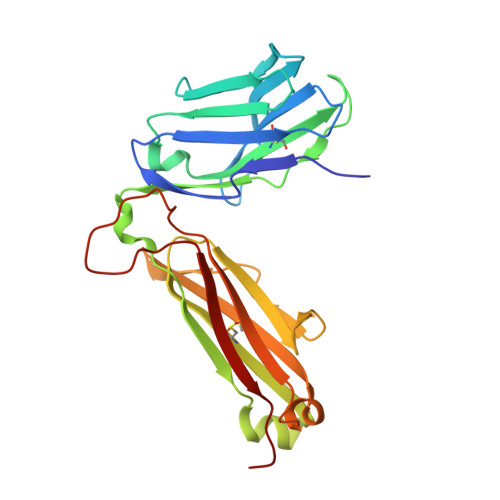
A conserved hydrophobic patch on Vbeta domains revealed by TCRbeta chain crystal structures: implications for pre-TCR dimerization
Zhou, B., Chen, Q., Mallis, R.J., Zhang, H., Liu, J.H., Reinherz, E.L., Wang, J.H.(2011) Front Immunol 2: 5-5
- PubMed: 22566796 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3389/fimmu.2011.00005
- Primary Citation Related Structures:
3Q5T, 3Q5Y - PubMed Abstract:
The αβ T cell receptor (TCR) is a multimeric complex whose β chain plays a crucial role in thymocyte development as well as antigen recognition by mature T lymphocytes. We report here crystal structures of individual β subunits, termed N15β (Vβ5.2Dβ2Jβ2.6Cβ2) and N30β (Vβ13Dβ1Jβ1.1Cβ2), derived from two αβ TCRs specific for the immunodominant vesicular stomatitis virus octapeptide (VSV-8) bound to the murine H-2K(b) MHC class I molecule. The crystal packing of the N15β structure reveals a homodimer formed through two Vβ domains. The Vβ/Vβ module is topologically very similar to the Vα/Vβ module in the N15αβ heterodimer. By contrast, in the N30β structure, the Vβ domain's external hydrophobic CFG face is covered by the neighboring molecule's Cβ domain. In conjunction with systematic investigation of previously published TCR single-subunit structures, we identified several conserved residues forming a concave hydrophobic patch at the center of the CFG outer face of the Vβ and other V-type Ig-like domains. This hydrophobic patch is shielded from solvent exposure in the crystal packing, implying that it is unlikely to be thermodynamically stable if exposed on the thymocyte surface. Accordingly, we propose a dimeric pre-TCR model distinct from those suggested previously by others and discuss its functional and structural implications.
- Laboratory of Immunobiology, Dana-Farber Cancer Institute, Harvard Medical School Boston, MA, USA.
Organizational Affiliation: