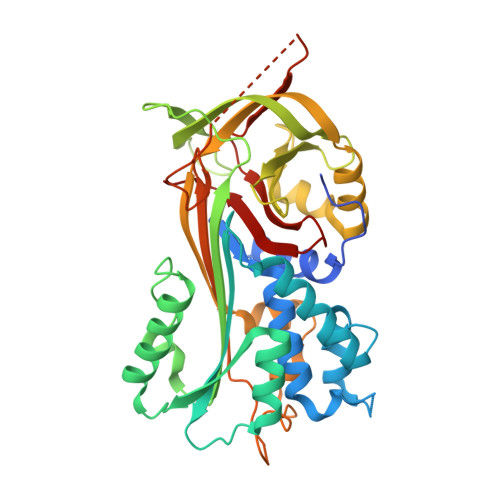
Crystal structure of native Anopheles gambiae serpin-2, a negative regulator of melanization in mosquitoes.
An, C., Lovell, S., Kanost, M.R., Battaile, K.P., Michel, K.(2011) Proteins 79: 1999-2003
- PubMed: 21465556 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/prot.23002
- Primary Citation Related Structures:
3PZF - PubMed Abstract:
Serpins are the dominant group of protease inhibitors in metazoans that control a wide variety of biological processes including major innate immune reactions. One of these inhibitors, SRPN2, controls melanization in mosquitoes – a powerful, arthropod-specific innate immune response. SRPN2 depletion from the hemolymph of adult female mosquitoes significantly reduces longevity and therefore this serpin is a potential target for novel insecticides. We report here the crystal structure of SRPN2 in its native conformation from the African malaria mosquito, Anopheles gambiae to 1.75 Å resolution. SRPN2 adopts a similar fold as observed for other serpins with a core of three β-sheets surrounded by nine α-helices with an exposed reactive center loop (RCL) that extends from the protein body. Similar to other native serpin structures, several residues within the reactive center loop were disordered and could not be modeled. Intriguingly, the N-terminal hinge of the RCL in SRPN2 was found to be inserted into β-sheet A, suggesting a potential activation mechanism analogous to heparin-mediated activation of Antithrombin III.
- Division of Biology, Kansas State University, Manhattan, Kansas 66506, USA.
Organizational Affiliation: