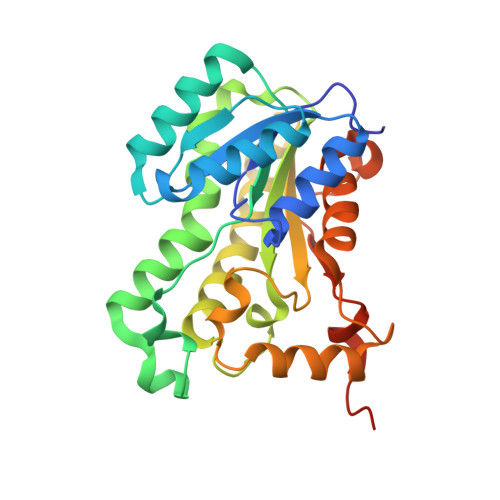
Structural basis of triclosan resistance
Jiten Singh, N., Shin, D.G., Lee, H.M., Kim, H.T., Chang, H.J., Cho, J.M., Kim, K.S., Ro, S.(2011) J Struct Biol 174: 173-179
- PubMed: 21094257 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2010.11.008
- Primary Citation Related Structures:
3PJD, 3PJE, 3PJF - PubMed Abstract:
Triclosan (5-chloro-2-(2,4-dichloro-phenoxy)-phenol, TCL) is a well known inhibitor against enoyl-acyl carrier protein reductase (ENR), an enzyme critical for cell-wall synthesis of bacteria. The inhibitory concentration at 50% inhibition (IC(50)) of TCL against the Escherichia coli ENR is 150nM for wild type (WT), 380, 470 and 68,500nM for Ala, Ser and Val mutants, respectively. To understand this high TCL resistance in the G93V mutant, we obtained the crystal structures of mutated ENRs complexed with TCL and NAD(+). The X-ray structural analysis along with the ab initio calculations and molecular dynamics simulations explains the serious consequence in the G93V mutant complex. The major interactions around TCL due to the aromatic(cation)-aromatic and hydrogen bonding interactions are found to be conserved both in WT and mutant complexes. Thus, the overall structural change of protein is minimal except that a flexible α-helical turn around TCL is slightly pushed away due to the presence of the bulky valine group. However, TCL shows substantial edge-to-face aromatic (π)-interactions with both the flexible R192-F203 region and the residues in the close vicinity of G93. The weakening of some edge-to-face aromatic interactions around TCL in the G93V mutant results in serious resistance to TCL. This understanding is beneficial to design new generation of antibiotics which will effectively act on the mutant ENRs.
- Center for Superfunctional Materials, Department of Chemistry, Pohang University of Science and Technology, Pohang 780-790, Republic of Korea.
Organizational Affiliation: