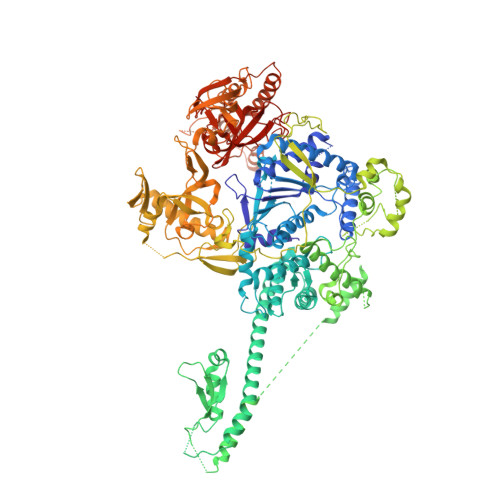
Structural and biochemical studies of the 5' -> 3' exoribonuclease Xrn1.
Chang, J.H., Xiang, S., Xiang, K., Manley, J.L., Tong, L.(2011) Nat Struct Mol Biol 18: 270-276
- PubMed: 21297639 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.1984
- Primary Citation Related Structures:
3PIE, 3PIF - PubMed Abstract:
The 5'→3' exoribonucleases (XRNs) have important functions in transcription, RNA metabolism and RNA interference. The structure of Rat1 (also known as Xrn2) showed that the two highly conserved regions of XRNs form a single, large domain that defines the active site of the enzyme. Xrn1 has a 510-residue segment after the conserved regions that is required for activity but is absent from Rat1/Xrn2. Here we report the crystal structures of Kluyveromyces lactis Xrn1 (residues 1-1,245, E178Q mutant), alone and in complex with a Mn(2+) ion in the active site. The 510-residue segment contains four domains (D1-D4), located far from the active site. Our mutagenesis and biochemical studies show that their functional importance results from their ability to stabilize the conformation of the N-terminal segment of Xrn1. These domains might also constitute a platform that interacts with protein partners of Xrn1.
- Department of Biological Sciences, Columbia University, New York, NY, USA.
Organizational Affiliation: