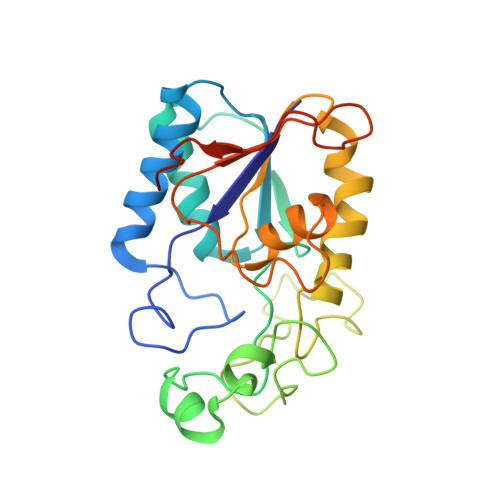
Structure and activity of phosphoglycerate mutase.
Winn, S.I., Watson, H.C., Harkins, R.N., Fothergill, L.A.(1981) Philos Trans R Soc London,ser B 293: 121-130
- PubMed: 6115412 Search on PubMed
- DOI: https://doi.org/10.1098/rstb.1981.0066
- Primary Citation Related Structures:
3PGM - PubMed Abstract:
The structure of yeast phosphoglycerate mutase determined by X-ray crystallographic and amino acid sequence studies has been interpreted in terms of the chemical, kinetic and mechanistic observations made on this enzyme. There are two histidine residues at the active site, with imidazole groups almost parallel to each other and approximately 0.4 nm apart, positioned close to the 2 and 3 positions of the substrate. The simplest interpretation of the available information suggests that a ping-pong type mechanism operates in which at least one of these histidine residues participates in the phosphoryl transfer reaction. The flexible C-terminal region also plays an important role in the enzymic reaction.