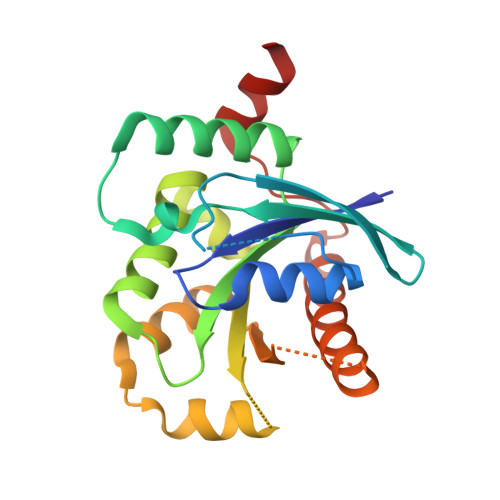
Crystal structure of human GTPase IMAP family member 2 in the nucleotide-free state
Shen, L., Tempel, W., Tong, Y., Guan, X., Nedyalkova, L., Wernimont, A.K., Mackenzie, F., Arrowsmith, C.H., Edwards, A.M., Bountra, C., Weigelt, J., Bochkarev, A., Andrews, D.W., Park, H.To be published.