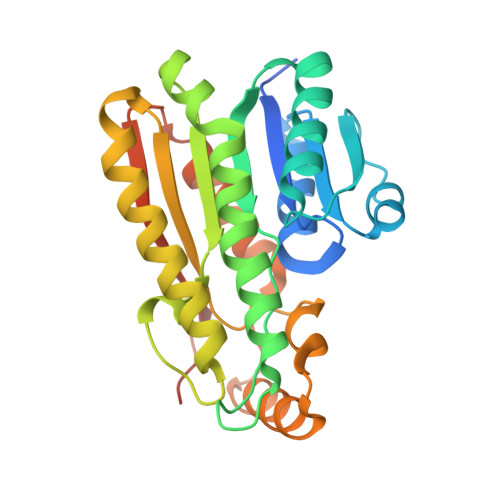
Structure of a NADPH-dependent blue fluorescent protein revealed the unique role of Gly176 on the fluorescence enhancement.
Kao, T.H., Chen, Y., Pai, C.H., Chang, M.C., Wang, A.H.J.(2011) J Struct Biol 174: 485-493
- PubMed: 21397029 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2011.02.010
- Primary Citation Related Structures:
3P19 - PubMed Abstract:
A NADPH-dependent blue fluorescent protein from Vibrio vulnificus CKM-1 (BFPvv) emits blue fluorescence under UV-exposure. Previously, the BFPvvD7 mutant generated by directed evolution displayed a fourfold enhancement in fluorescent intensity. Herein, a further increase in fluorescence in the new BFPvvD8 mutant, with three additional mutations from BFPvvD7, was made. To understand the underlying mechanism of the increased fluorescent intensity of BFPvv, we solved the BFPvvD8-NADPH complex structure. Accompanied with lifetime detection, we proposed that the enhanced intensity is related to the conformational change caused by a glycine residue (Gly176) mutated to other non-glycine residues at a turn close to the NADPH binding site. We also observed the Förster resonance energy transfer (FRET) from our BFPvvD8 to each of the GFP-like fluorescent proteins, mTFP1 and EGFP, joined by an eight-residue linker between the N-terminal of BFPvvD8 and the C-terminal of GFPs. Taken together, with the newly solved BFPvvD8 structure, our results not only provide new considerations within the rational-based protein engineering of this NADPH-dependent BFP, but also suggest that BFPvvD8 could be a potential candidate in FRET-based biosensor techniques.
- Institute of Biological Chemistry, Academia Sinica, Taipei, Taiwan.
Organizational Affiliation: