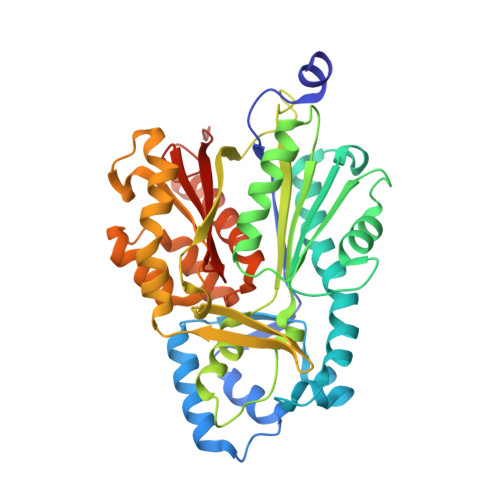
A hydrophobic cavity discovered in a curcumin synthase facilitates utilization of a beta-keto acid as an extender substrate for the atypical type III polyleteide synthase
Katsuyama, Y., Miyazono, K., Tanokura, M., Ohnishi, Y., Horinouchi, S.To be published.