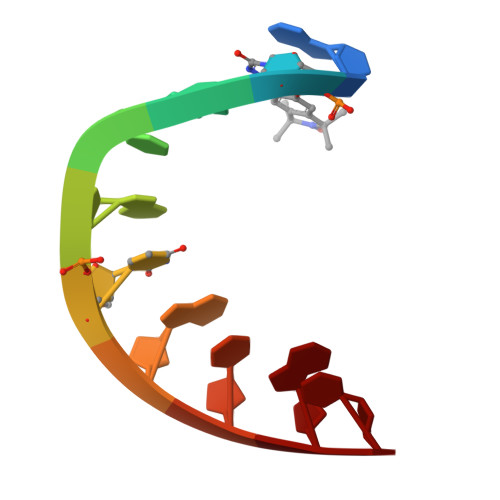
Crystal structure of a DNA containing the planar, phenoxazine-derived bi-functional spectroscopic probe C.
Edwards, T.E., Cekan, P., Reginsson, G.W., Shelke, S.A., Ferre-D'Amare, A.R., Schiemann, O., Sigurdsson, S.T.(2011) Nucleic Acids Res 10: 4419-4426
- PubMed: 21252294 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkr015
- Primary Citation Related Structures:
3OT0 - PubMed Abstract:
Previously, we developed the deoxycytosine analog Ç (C-spin) as a bi-functional spectroscopic probe for the study of nucleic acid structure and dynamics using electron paramagnetic resonance (EPR) and fluorescence spectroscopy. To understand the effect of Ç on nucleic acid structure, we undertook a detailed crystallographic analysis. A 1.7 Å resolution crystal structure of Ç within a decamer duplex A-form DNA confirmed that Ç forms a non-perturbing base pair with deoxyguanosine, as designed. In the context of double-stranded DNA Ç adopted a planar conformation. In contrast, a crystal structure of the free spin-labeled base ç displayed a ∼ 20° bend at the oxazine linkage. Density function theory calculations revealed that the bent and planar conformations are close in energy and exhibit the same frequency for bending. These results indicate a small degree of flexibility around the oxazine linkage, which may be a consequence of the antiaromaticity of a 16-π electron ring system. Within DNA, the amplitude of the bending motion is restricted, presumably due to base-stacking interactions. This structural analysis shows that the Ç forms a planar, structurally non-perturbing base pair with G indicating it can be used with high confidence in EPR- or fluorescence-based structural and dynamics studies.
- Emerald BioStructures, Bainbridge Island, WA 98110, USA.
Organizational Affiliation: