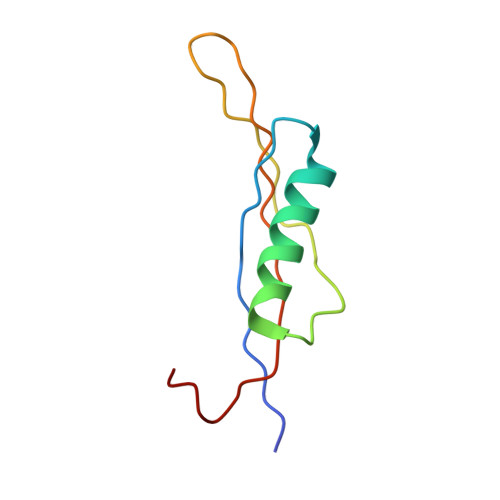
Crystal structure of calcium dodecin (Rv0379), from Mycobacterium tuberculosis with a unique calcium-binding site.
Arockiasamy, A., Aggarwal, A., Savva, C.G., Holzenburg, A., Sacchettini, J.C.(2011) Protein Sci 20: 827-833
- PubMed: 21370306 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.607
- Primary Citation Related Structures:
3ONR - PubMed Abstract:
In eukaryotes, calcium-binding proteins play a pivotal role in diverse cellular processes, and recent findings suggest similar roles for bacterial proteins at different stages in their life cycle. Here, we report the crystal structure of calcium dodecin, Rv0379, from Mycobacterium tuberculosis with a dodecameric oligomeric assembly and a unique calcium-binding motif. Structure and sequence analysis were used to identify orthologs of Rv0379 with different ligand-binding specificity.
- Department of Biochemistry and Biophysics, Texas A&M University, College Station, Texas 77843-2128, USA.
Organizational Affiliation: