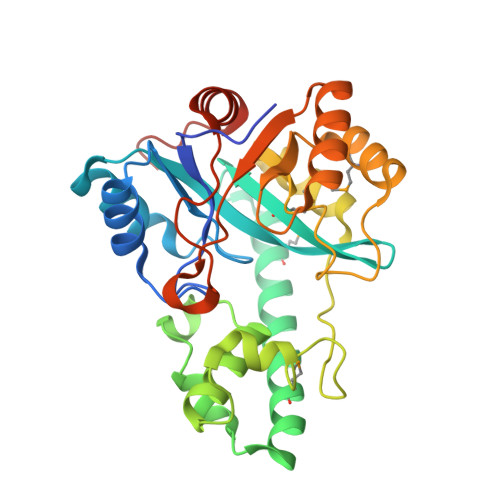
Crystal structure of a metal-dependent phosphoesterase (YP_910028.1) from Bifidobacterium adolescentis: Computational prediction and experimental validation of phosphoesterase activity.
Han, G.W., Ko, J., Farr, C.L., Deller, M.C., Xu, Q., Chiu, H.J., Miller, M.D., Sefcikova, J., Somarowthu, S., Beuning, P.J., Elsliger, M.A., Deacon, A.M., Godzik, A., Lesley, S.A., Wilson, I.A., Ondrechen, M.J.(2011) Proteins 79: 2146-2160
- PubMed: 21538547 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/prot.23035
- Primary Citation Related Structures:
3E0F, 3O0F - PubMed Abstract:
The crystal structures of an unliganded and adenosine 5'-monophosphate (AMP) bound, metal-dependent phosphoesterase (YP_910028.1) from Bifidobacterium adolescentis are reported at 2.4 and 1.94 Å, respectively. Functional characterization of this enzyme was guided by computational analysis and then confirmed by experiment. The structure consists of a polymerase and histidinol phosphatase (PHP, Pfam: PF02811) domain with a second domain (residues 105-178) inserted in the middle of the PHP sequence. The insert domain functions in binding AMP, but the precise function and substrate specificity of this domain are unknown. Initial bioinformatics analyses yielded multiple potential functional leads, with most of them suggesting DNA polymerase or DNA replication activity. Phylogenetic analysis indicated a potential DNA polymerase function that was somewhat supported by global structural comparisons identifying the closest structural match to the alpha subunit of DNA polymerase III. However, several other functional predictions, including phosphoesterase, could not be excluded. Theoretical microscopic anomalous titration curve shapes, a computational method for the prediction of active sites from protein 3D structures, identified potential reactive residues in YP_910028.1. Further analysis of the predicted active site and local comparison with its closest structure matches strongly suggested phosphoesterase activity, which was confirmed experimentally. Primer extension assays on both normal and mismatched DNA show neither extension nor degradation and provide evidence that YP_910028.1 has neither DNA polymerase activity nor DNA-proofreading activity. These results suggest that many of the sequence neighbors previously annotated as having DNA polymerase activity may actually be misannotated.
- Joint Center for Structural Genomics, Scripps Research Institute, La Jolla, California 92037, USA.
Organizational Affiliation: