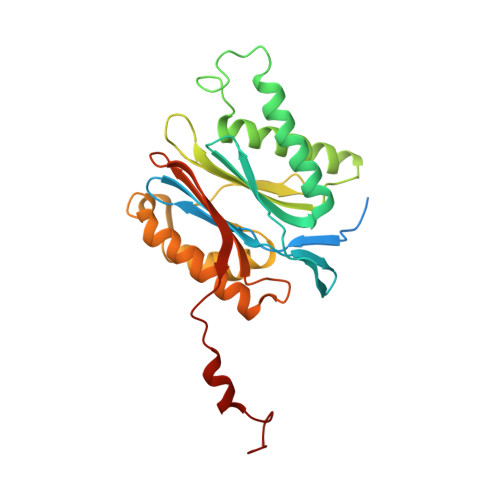
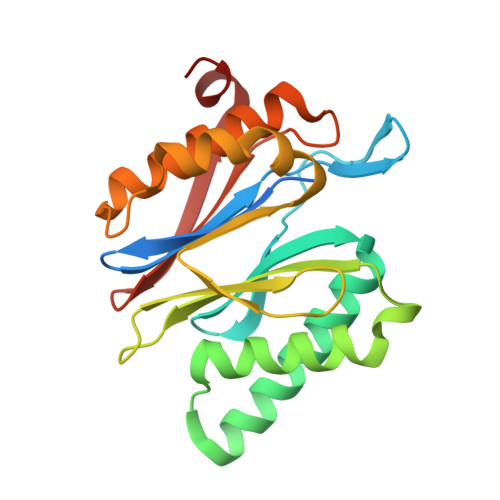
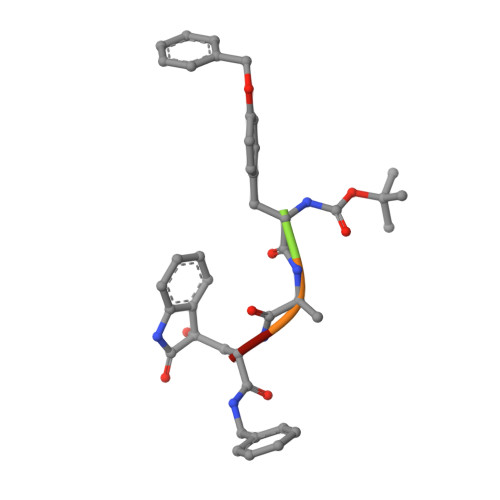
20S proteasome inhibition: designing noncovalent linear peptide mimics of the natural product TMC-95A.
Groll, M., Gallastegui, N., Marechal, X., Le Ravalec, V., Basse, N., Richy, N., Genin, E., Huber, R., Moroder, L., Vidal, J., Reboud-Ravaux, M.(2010) ChemMedChem 5: 1701-1705
- PubMed: 20715286 Search on PubMed
- DOI: https://doi.org/10.1002/cmdc.201000293
- Primary Citation Related Structures:
3NZJ, 3NZW, 3NZX - Department Chemie, Lehrstuhl für Biochemie, Technische Universität München, Garching Germany. michael.groll@ch.tum.de
Organizational Affiliation: